DNA replication in eukaryotes ensures every cell gets an accurate copy of the genome before division. Because eukaryotic genomes are large and packaged into chromatin, the process requires multiple origins of replication, several specialized polymerases, and tight coordination with the cell cycle.
DNA Replication in Eukaryotes
Process of eukaryotic DNA replication
Eukaryotic replication follows a series of tightly controlled stages. Here's how the process unfolds, from start to finish.
1. Initiation
Replication begins at origins of replication, specific DNA sequences scattered along each chromosome. Eukaryotic cells have thousands of these origins so the entire genome can be copied in a reasonable time frame.
Before replication can start, a pre-replication complex (pre-RC) must assemble at each origin:
- The origin recognition complex (ORC) binds to the origin sequence.
- Cdc6 and Cdt1 proteins then recruit the MCM helicase complex to the origin.
- Licensing factors ensure each origin fires only once per cell cycle, preventing the same stretch of DNA from being copied twice.
The MCM helicase stays inactive until it's phosphorylated by two kinases: Dbf4-dependent kinase (DDK) and cyclin-dependent kinase (CDK). This activation step links replication directly to cell cycle progression.
2. Unwinding and strand separation
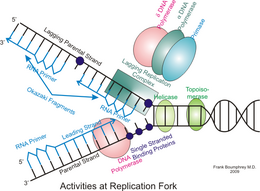
Once activated, the MCM helicase unwinds the double helix, creating a replication fork where the two strands separate. RPA (Replication Protein A), a single-stranded DNA binding protein, coats the exposed single strands to keep them from snapping back together or being degraded.
3. Priming and elongation
DNA polymerases can't start a new strand from scratch. They need a short RNA primer laid down by primase. From there, synthesis proceeds differently on each strand:
- Leading strand: DNA polymerase ε (epsilon) synthesizes DNA continuously in the 5' to 3' direction, following the replication fork as it opens.
- Lagging strand: DNA polymerase δ (delta) synthesizes DNA in short segments called Okazaki fragments (each about 100–200 nucleotides in eukaryotes). Because this strand runs in the opposite orientation, the polymerase has to repeatedly start new fragments as more template is exposed.
DNA polymerase α (alpha) plays a special role early on: it extends each RNA primer by adding a short stretch of DNA, then hands off to polymerase ε or δ for the bulk of synthesis.
After the Okazaki fragments are made, the RNA primers are removed and replaced with DNA by polymerase δ. DNA ligase then seals the remaining nicks to create a continuous strand.
4. Termination
Replication forks from neighboring origins eventually converge and meet. Topoisomerases resolve supercoils and tangles that build up ahead of and between converging forks. The newly synthesized DNA is then proofread; mismatch repair enzymes correct errors that the polymerases' own proofreading missed.

Function of telomeres and telomerase
Linear chromosomes create a unique problem: conventional DNA polymerases can't fully replicate the very end of each chromosome. This is called the end-replication problem, and it means a small amount of DNA is lost from chromosome tips with every round of replication.
Telomeres are the solution. These are repetitive, non-coding DNA sequences at chromosome ends. In humans, the repeat unit is TTAGGG, present in thousands of copies. Telomeres act as a disposable buffer so that the DNA lost each cycle doesn't eat into important genes. They also prevent chromosome ends from being mistaken for broken DNA and fusing together.
Telomerase is the enzyme that counteracts telomere shortening. It's a ribonucleoprotein with two key components:
- TERC (Telomerase RNA Component): carries a built-in RNA template complementary to the telomere repeat sequence.
- TERT (Telomerase Reverse Transcriptase): uses that RNA template to add new TTAGGG repeats onto the 3' end of the chromosome.
Telomerase is active in stem cells and germ cells (which need to divide indefinitely) but is repressed in most somatic cells. This matters for aging and disease:
- As somatic cells divide, telomeres gradually shorten. When they reach a critically short length, the cell enters senescence (permanent cell cycle arrest) or undergoes apoptosis (programmed cell death).
- Many cancer cells reactivate telomerase, which allows them to maintain telomere length and divide without limit. This is one reason cancers can grow indefinitely.
DNA replication in prokaryotes vs eukaryotes
Several key differences reflect the greater size and complexity of eukaryotic genomes:
| Feature | Prokaryotes | Eukaryotes |
|---|---|---|
| Replication rate | ~1,000 nucleotides/sec (E. coli) | ~50 nucleotides/sec (humans) |
| Genome size | Smaller (~4.6 million bp in E. coli) | Much larger (~3 billion bp in humans) |
| Origins of replication | Single origin per circular chromosome | Thousands of origins along linear chromosomes |
| Replication machinery | Simpler (DNA Pol III, helicase, primase) | More complex (multiple polymerases, ORC, MCM, licensing factors) |
| Cell cycle regulation | No distinct G1/G2 phases; replication is continuous | Replication restricted to S phase of a tightly regulated cell cycle |
Eukaryotes compensate for their slower polymerases by firing many origins simultaneously. Even though each fork moves more slowly, the whole genome gets copied within a few hours.
Key features of eukaryotic DNA replication
- Semiconservative: Each daughter molecule contains one original strand and one newly synthesized strand. This was demonstrated by the Meselson-Stahl experiment.
- Bidirectional: Two replication forks move in opposite directions from each origin, doubling the speed at which a region is copied.
- Chromatin remodeling required: DNA in eukaryotes is wrapped around histone proteins. Chromatin must be loosened ahead of the replication fork and reassembled behind it so the new DNA is properly packaged.
- High processivity: Once DNA polymerases (aided by the sliding clamp PCNA) latch onto the template, they can synthesize long stretches of DNA without falling off.