Basics of Monte Carlo integration
Monte Carlo integration uses random sampling to estimate integrals that are too complex to solve analytically. In Bayesian statistics, this matters constantly: posterior distributions, marginal likelihoods, and model evidence all involve integrals that rarely have closed-form solutions. Monte Carlo methods let you approximate these integrals numerically, with quantifiable uncertainty around the estimates.
Definition and purpose
The core idea is straightforward: instead of solving an integral exactly, you draw random samples from the integration domain and average the function values at those points. This average converges to the true integral as you increase the number of samples.
Monte Carlo integration is especially valuable when:
- The integral is high-dimensional (more than a few parameters)
- The probability distribution has a complicated shape
- Deterministic numerical methods (like quadrature) become impractical
Every Monte Carlo estimate comes with a measure of uncertainty, which fits naturally into the Bayesian framework where quantifying uncertainty is the whole point.
Historical background
Stanislaw Ulam and John von Neumann developed Monte Carlo methods during the Manhattan Project in the 1940s. They needed to simulate neutron diffusion and nuclear fission processes, problems involving so many random interactions that analytical solutions were hopeless.
The name comes from the Monte Carlo Casino in Monaco, a nod to the role of randomness in the method. As computing power grew through the late 20th century, Monte Carlo methods spread into physics, finance, computer graphics, and eventually became central to Bayesian computation.
Relation to Bayesian statistics
Bayesian inference requires computing integrals over parameter spaces. For example, the posterior mean of a parameter is:
For anything beyond simple conjugate models, this integral has no closed form. Monte Carlo integration makes Bayesian inference practical by:
- Approximating posterior distributions through random samples
- Estimating marginal likelihoods for model comparison
- Handling high-dimensional parameter spaces where grid-based methods fail
- Enabling Bayes factors and hypothesis testing in complex models
Principles of Monte Carlo methods
Monte Carlo methods rest on a few key ideas from probability theory. Understanding these foundations helps you reason about why Monte Carlo estimates work and when you can trust them.
Random sampling
Everything starts with generating random numbers from a specified probability distribution. In practice, computers use pseudo-random number generators (PRNGs) like the Mersenne Twister algorithm, which produce sequences that behave like truly random draws for statistical purposes.
Setting a seed for your PRNG is important for reproducibility. Anyone who runs your code with the same seed gets the same "random" samples and the same results.
Several techniques exist for converting uniform random numbers into draws from other distributions:
- Inverse transform sampling: Apply the inverse CDF of the target distribution to uniform samples
- Rejection sampling: Propose samples from a simpler distribution and accept/reject them based on the target density
- Box-Muller transform: Converts pairs of uniform samples into standard normal variates
Law of large numbers
The law of large numbers (LLN) is the theoretical backbone of Monte Carlo integration. It states that the sample mean converges to the true expected value as the number of samples grows:
The weak LLN guarantees convergence in probability; the strong LLN guarantees almost sure convergence. For Monte Carlo purposes, the practical takeaway is the same: draw enough samples, and your estimate will be close to the true value.
Central limit theorem
The central limit theorem (CLT) tells you how fast convergence happens and lets you quantify uncertainty. Regardless of the shape of the underlying distribution (as long as it has finite variance), the distribution of the sample mean approaches a normal distribution as increases.
This means you can construct confidence intervals around your Monte Carlo estimates. If your estimate is with standard error , then an approximate 95% confidence interval is . The CLT also justifies normal approximations that show up frequently in large-sample Bayesian inference.
Monte Carlo integration techniques
Different problems call for different sampling strategies. The techniques below range from the simplest baseline to more sophisticated approaches that reduce variance and improve efficiency.
Simple Monte Carlo integration
The most basic version: draw random samples from a distribution , then estimate the integral as:
The convergence rate is . That means to cut your error in half, you need four times as many samples. This rate holds regardless of the dimensionality of the problem, which is a major advantage over deterministic quadrature methods in high dimensions.
Simple Monte Carlo works well for low-dimensional problems with straightforward integration domains, but it can be inefficient when the integrand varies a lot across the domain.
Importance sampling
When the integrand is concentrated in a small region of the domain, most simple Monte Carlo samples contribute very little to the estimate. Importance sampling fixes this by drawing samples from a proposal distribution that concentrates samples where the integrand is large.
The corrected estimate is:
The ratio is called the importance weight. It corrects for the fact that you're sampling from instead of .
The catch: your choice of matters enormously. A good proposal distribution (one that closely matches the shape of ) can dramatically reduce variance. A poor choice can make things worse than simple Monte Carlo, especially if the importance weights have heavy tails.
Stratified sampling
Stratified sampling divides the integration domain into non-overlapping regions (strata) and draws samples independently within each one. The estimate combines results from all strata:
where is the weight (typically the volume) of stratum and is the number of samples in that stratum.
This reduces variance by ensuring every region of the domain gets represented, which prevents the "clumping" that can happen with purely random sampling. It's especially effective when the integrand varies a lot in certain regions. You can also combine stratified sampling with importance sampling for further gains.
Applications in Bayesian inference
Posterior distribution estimation
The most common application: you want to know the posterior distribution but can't compute it analytically. Monte Carlo methods generate samples from the posterior, and you use those samples to estimate whatever you need:
- Posterior mean: average the samples
- Credible intervals: sort the samples and take quantiles (e.g., the 2.5th and 97.5th percentiles for a 95% interval)
- Posterior predictive distribution: for each sampled , simulate new data, giving you a distribution of predictions that accounts for parameter uncertainty
In practice, MCMC methods (covered later in this unit) are the most common way to generate these posterior samples.
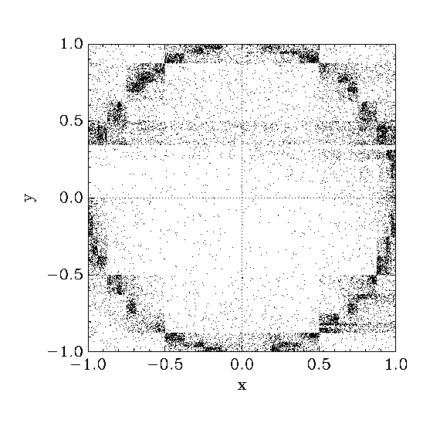
Marginal likelihood calculation
The marginal likelihood (also called the evidence) is:
This integral over the entire parameter space is needed for Bayesian model comparison but is notoriously difficult to compute. Several Monte Carlo approaches exist:
- Harmonic mean estimator: Simple to compute from posterior samples, but numerically unstable and often unreliable
- Bridge sampling: Uses samples from both the prior and posterior to construct a more stable estimate
- Nested sampling: Transforms the multi-dimensional integral into a one-dimensional problem by sampling in nested shells of increasing likelihood
Model comparison
With marginal likelihoods in hand, you can compute Bayes factors to compare competing models. The Bayes factor for model versus is the ratio of their marginal likelihoods:
This naturally penalizes overly complex models because the marginal likelihood integrates over the full prior, spreading probability mass across the parameter space. More specialized approaches like Reversible Jump MCMC can simultaneously explore different model structures and estimate posterior model probabilities.
Advantages and limitations
Computational efficiency
Monte Carlo methods scale much better to high dimensions than deterministic integration methods like quadrature, which require an exponentially growing number of evaluation points as dimensions increase. Monte Carlo's convergence rate of is independent of dimensionality.
Additional efficiency gains come from:
- Parallelization (many samples can be drawn independently)
- Adaptive algorithms that focus computational effort on important regions
- The ability to handle irregular or complex integration domains without special treatment
Accuracy vs. sample size
The standard Monte Carlo error decreases as . Concretely, if 1,000 samples give you a standard error of 0.1, you'd need 100,000 samples to get the standard error down to 0.01. This means brute-force accuracy improvements are expensive.
Variance reduction techniques (importance sampling, stratified sampling, control variates) can effectively give you the accuracy of a much larger sample at lower computational cost. Monitoring convergence diagnostics is essential so you know when your estimates are reliable enough to stop sampling.
Curse of dimensionality
While Monte Carlo methods handle high dimensions better than grid-based methods, they aren't immune to dimensionality challenges. As the number of parameters grows, the volume of the parameter space explodes, and most of that volume contains negligible posterior probability.
Simple Monte Carlo and importance sampling struggle in very high dimensions because it becomes nearly impossible to find a good proposal distribution. This is precisely why MCMC methods (Metropolis-Hastings, Hamiltonian Monte Carlo) and other specialized techniques were developed: they explore high-dimensional spaces by taking correlated steps through regions of high probability rather than sampling the whole space at once.
Implementation in practice
Algorithm design
Choosing the right Monte Carlo technique depends on your problem:
- Assess dimensionality: Low-dimensional problems (fewer than ~5 parameters) may work fine with simple Monte Carlo or importance sampling. Higher dimensions typically need MCMC.
- Examine the posterior shape: Multimodal posteriors require methods that can jump between modes (e.g., parallel tempering, sequential Monte Carlo).
- Apply variance reduction where possible: If you have analytical knowledge about part of the integrand, use control variates or Rao-Blackwellization.
- Incorporate domain knowledge: Informative proposal distributions based on what you know about the problem can dramatically improve efficiency.
Software tools and libraries
Several mature software packages handle Monte Carlo integration and Bayesian inference:
- Stan (via RStan, PyStan, CmdStan): Implements Hamiltonian Monte Carlo; widely used for complex Bayesian models
- PyMC (formerly PyMC3): Python-based probabilistic programming with NUTS sampler
- BUGS/JAGS: Gibbs sampling frameworks; good for conditionally conjugate models
- TensorFlow Probability: Scalable Bayesian inference with GPU support
- SciPy/NumPy: Lower-level tools for implementing custom Monte Carlo algorithms in Python
When choosing a tool, consider the model complexity you need, the sampling algorithms available, and the size of the user community (which affects documentation quality and troubleshooting support).
Parallel computing considerations
Many Monte Carlo algorithms are embarrassingly parallel, meaning independent samples can be generated simultaneously on different processors with no communication between them. This makes them natural fits for multi-core CPUs and GPUs.
For MCMC, parallelism is trickier because each sample depends on the previous one. Common strategies include running multiple independent chains in parallel and using ensemble samplers. Be mindful of communication overhead in distributed computing setups, and use libraries designed for parallel Monte Carlo (e.g., Dask, Ray, or the built-in parallelism in Stan).
Advanced Monte Carlo techniques
Markov Chain Monte Carlo (MCMC)
MCMC constructs a Markov chain whose stationary distribution is the target posterior. After a burn-in period, samples from the chain approximate draws from the posterior. This is the topic of the rest of Unit 7, but the key algorithms are:
- Metropolis-Hastings: Proposes a new point and accepts/rejects it based on the ratio of posterior densities. The foundation for most MCMC methods.
- Gibbs sampling: Samples each parameter in turn from its full conditional distribution. Efficient when those conditionals are available in closed form.
- Hamiltonian Monte Carlo (HMC): Uses gradient information to propose distant points with high acceptance probability, making it much more efficient in high dimensions.
- Adaptive MCMC: Automatically tunes the proposal distribution during sampling to improve mixing.
Sequential Monte Carlo
Also called particle filtering, sequential Monte Carlo (SMC) maintains a population of weighted samples ("particles") that evolve as new data arrives. This makes SMC particularly suited for:
- Dynamic models where the posterior updates over time
- Non-linear and non-Gaussian state-space models
- Applications in tracking, robotics, and time series analysis
SMC can also be used for static problems by introducing a sequence of intermediate distributions that gradually transition from the prior to the posterior.
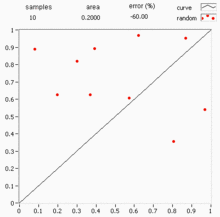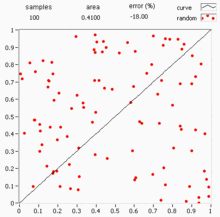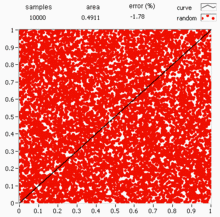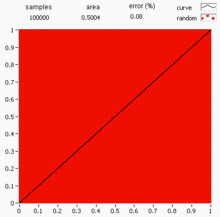
Quasi-Monte Carlo methods
Standard Monte Carlo uses pseudo-random numbers, but quasi-Monte Carlo (QMC) replaces them with low-discrepancy sequences (e.g., Sobol or Halton sequences) that fill the space more evenly. This can improve the convergence rate from to or better for sufficiently smooth integrands.
QMC works best in low to moderate dimensions. Scrambled quasi-Monte Carlo randomizes the low-discrepancy sequences, combining the better coverage of QMC with the ability to estimate errors statistically. QMC has seen significant use in financial modeling and computer graphics.
Error estimation and convergence
Monte Carlo standard error
The Monte Carlo standard error (MCSE) tells you how precise your estimate is. For simple Monte Carlo integration:
You estimate from your samples. For importance sampling, the formula adjusts to account for the variability of the importance weights.
For MCMC, samples are correlated, so the naive formula underestimates the true standard error. The batch means method handles this by dividing the chain into batches, computing the mean of each batch, and using the variability across batch means to estimate the standard error. The effective sample size (ESS) accounts for autocorrelation: if your chain has high autocorrelation, the ESS will be much smaller than the actual number of samples.
Convergence diagnostics
For MCMC specifically, you need to verify that the chain has reached its stationary distribution before trusting the samples. Common diagnostics include:
- Gelman-Rubin statistic (): Run multiple chains from different starting points and compare within-chain variance to between-chain variance. Values close to 1.0 (typically below 1.01) suggest convergence.
- Geweke test: Compares the mean of the first portion of a chain to the mean of the last portion. A significant difference suggests the chain hasn't converged.
- Trace plots: Visual inspection of parameter values over iterations. A well-mixed chain looks like a "fuzzy caterpillar" with no trends or long excursions.
- Autocorrelation plots: Show how correlated successive samples are. High autocorrelation means the chain is exploring slowly.
Variance reduction techniques
These methods squeeze more accuracy out of the same number of samples:
- Antithetic variates: For each random sample, also evaluate the function at a "mirror" sample. The negative correlation between the two reduces overall variance.
- Control variates: If you know the expected value of a related function , you can subtract a correction term from your estimator to reduce variance.
- Rao-Blackwellization: Replace a Monte Carlo average with a conditional expectation wherever possible. This always reduces variance (or leaves it unchanged) and is especially useful in Gibbs sampling.
Case studies and examples
Bayesian parameter estimation
Consider estimating the coefficients of a logistic regression model for medical diagnosis (e.g., predicting disease presence from biomarkers). The posterior distribution over regression coefficients has no closed form.
Using MCMC, you draw thousands of samples from the posterior. Each sample is a complete set of regression coefficients. From these samples, you can compute posterior means, 95% credible intervals for each coefficient, and posterior predictive probabilities for new patients. Comparing these results with maximum likelihood estimates reveals how prior information and uncertainty quantification change the conclusions.
Hierarchical models
A hierarchical model for student performance across multiple schools estimates both school-level effects and an overall distribution of effects. Gibbs sampling works well here because the conditional distributions at each level are often available in closed form.
The key Bayesian feature is partial pooling: schools with little data get "shrunk" toward the overall mean, borrowing strength from other schools. Monte Carlo samples from the posterior let you visualize this shrinkage and quantify uncertainty at both the school and population levels.
High-dimensional integrals
In astrophysics, models for phenomena like gravitational wave signals can have dozens of parameters. Computing the marginal likelihood for model comparison requires integrating over all of them.
Nested sampling is often the method of choice here. It handles multimodal posteriors and simultaneously produces both posterior samples and a marginal likelihood estimate. Comparing nested sampling's efficiency against other approaches (like thermodynamic integration) on these problems illustrates the practical trade-offs between different Monte Carlo strategies.
Challenges and future directions
Scalability issues
As datasets grow to millions of observations, evaluating the likelihood at every MCMC step becomes a bottleneck. Subsampling methods (like stochastic gradient MCMC) use random mini-batches of data to approximate the likelihood, trading some accuracy for massive speed gains. Distributed computing frameworks and streaming algorithms are active areas of research for scaling Bayesian Monte Carlo to big data settings.
Adaptive Monte Carlo methods
Adaptive methods automatically tune their own parameters during sampling. Examples include adaptive Metropolis algorithms that learn the proposal covariance from past samples, and self-tuning importance sampling schemes. The theoretical challenge is proving that these adaptations don't break the convergence guarantees that make Monte Carlo methods valid.
Integration with machine learning
Recent work combines Monte Carlo methods with deep learning. Normalizing flows and other neural network architectures can learn flexible proposal distributions for MCMC or importance sampling. Variational inference offers a faster (but approximate) alternative to MCMC, and hybrid methods try to get the best of both worlds. Bayesian deep learning uses Monte Carlo techniques to quantify uncertainty in neural network predictions, which matters for safety-critical applications like medical imaging and autonomous driving.