RNA is the single-stranded counterpart to DNA, but its roles extend far beyond simply carrying genetic information. Because RNA can fold back on itself and form complex three-dimensional shapes, it takes on structural, catalytic, and regulatory functions that DNA cannot. This topic covers RNA's structural features, the major types involved in protein synthesis, and the regulatory RNAs that fine-tune gene expression.
RNA Structure
Single-Stranded Structure and Hairpin Loops
RNA is built from a single strand of nucleotides, which immediately distinguishes it from double-stranded DNA. This single-stranded nature is what gives RNA its functional versatility: regions within the same strand can fold back and base-pair with each other, creating secondary structures.
The most common of these is the hairpin loop (also called a stem-loop). A hairpin forms when a stretch of nucleotides is complementary to another stretch further along the same molecule. The complementary regions pair up to form the "stem," while the unpaired nucleotides in between loop out at the top. These structures are critical for function. For example, tRNA folds into a cloverleaf shape held together by multiple stem-loops, and ribozymes depend on specific hairpin arrangements for catalytic activity.
Base pairing in RNA follows Watson-Crick rules, but with one key difference from DNA: adenine (A) pairs with uracil (U) instead of thymine, and guanine (G) pairs with cytosine (C). RNA can also form non-canonical base pairs (like G-U wobble pairs), which adds further structural diversity.
Codons and Anticodons in Genetic Information Transfer
A codon is a three-nucleotide sequence in mRNA that specifies a particular amino acid during translation. With four possible bases at each of three positions, there are possible codons. Of these:
- AUG serves as the universal start codon (and codes for methionine)
- UAA, UAG, and UGA are stop codons that signal the end of translation
An anticodon is the complementary three-nucleotide sequence on a tRNA molecule. During translation, the anticodon on a tRNA base-pairs with the codon on the mRNA, ensuring the correct amino acid is added to the growing polypeptide chain.
Wobble base pairing is worth understanding here. The third position of the codon (and first position of the anticodon) can tolerate non-standard pairing. This means a single tRNA can sometimes recognize more than one codon. Wobble pairing explains why organisms can get by with fewer than 61 different tRNAs despite having 61 sense codons.
RNA Modifications: 5' Cap and 3' Poly-A Tail
Eukaryotic mRNA undergoes two major modifications at its ends before leaving the nucleus:
5' Cap:
- A modified guanine nucleotide (7-methylguanosine) is added to the 5' end of the mRNA via an unusual 5'-to-5' triphosphate linkage
- Functions: protects mRNA from degradation by exonucleases, helps recruit the ribosome for translation initiation, and facilitates nuclear export
3' Poly-A Tail:
- A string of 50 to 250 adenine nucleotides is added to the 3' end by the enzyme poly-A polymerase (this sequence is not encoded in the DNA template)
- Functions: stabilizes the mRNA, aids in nuclear export, and promotes translation efficiency
Together, these two modifications act as molecular shields. As the mRNA ages, the poly-A tail gradually shortens. Once it's too short, the mRNA becomes vulnerable to degradation. This is one way cells control how long an mRNA molecule remains active.
Types of RNA
Messenger RNA (mRNA) Structure and Function
mRNA carries the genetic instructions from DNA in the nucleus to ribosomes in the cytoplasm, where those instructions are translated into protein. It's transcribed from the template (antisense) strand of DNA.
In eukaryotes, the initial transcript (pre-mRNA) undergoes several processing steps before export:
- Addition of the 5' cap
- Addition of the 3' poly-A tail
- Splicing to remove introns (non-coding sequences), leaving only exons joined together
Mature mRNA also contains untranslated regions (UTRs) at both the 5' and 3' ends. These UTRs don't code for protein but play important roles in regulating translation efficiency, mRNA localization, and stability.
mRNA lifespan varies considerably. In prokaryotes, mRNAs typically last minutes, allowing rapid responses to environmental changes. In eukaryotes, mRNAs can persist for hours to days, depending on the gene and cell type.
Transfer RNA (tRNA) and Its Role in Translation
tRNA molecules are the adaptors of translation. Each tRNA carries a specific amino acid to the ribosome and matches it to the correct mRNA codon.
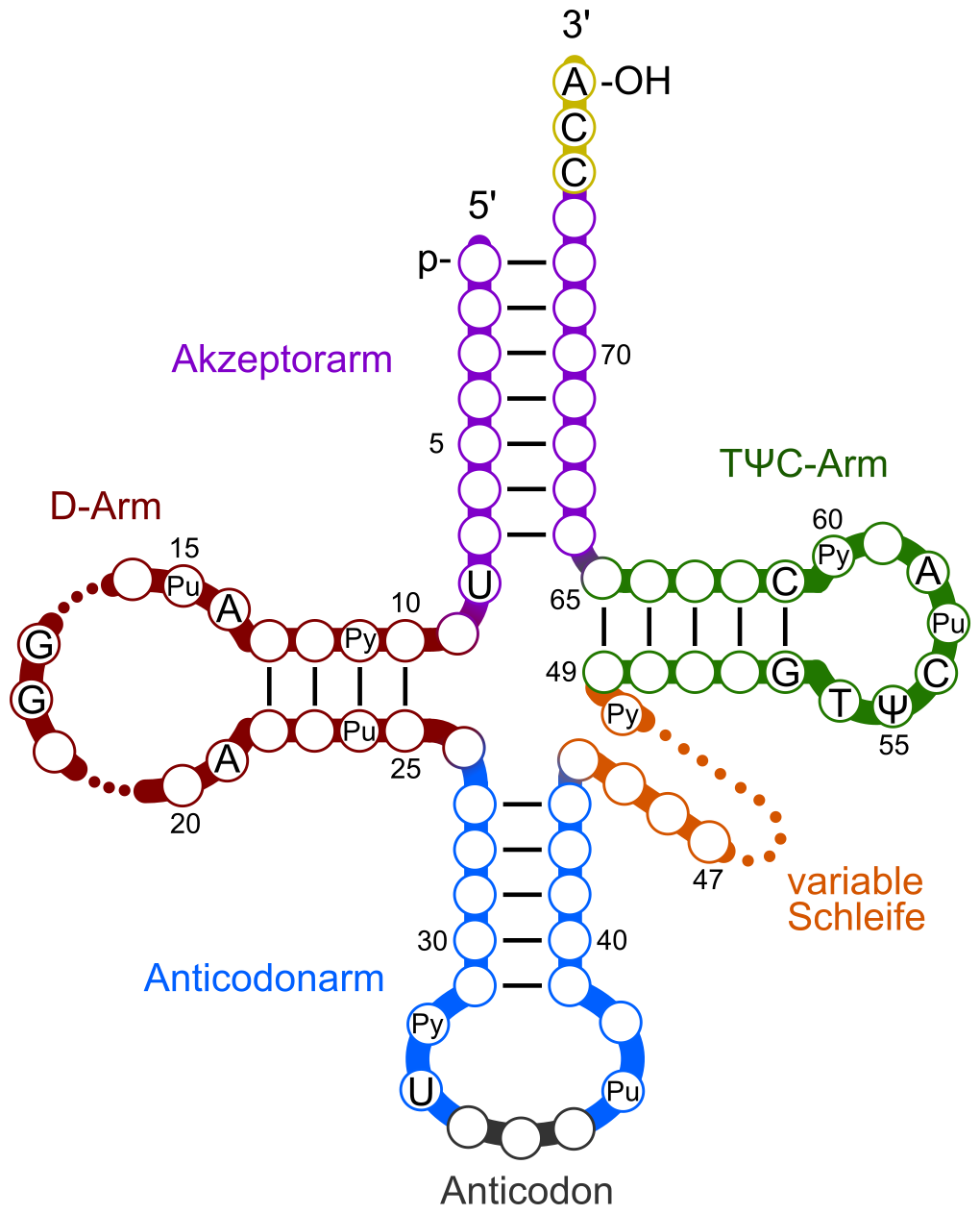
Key structural features of tRNA:
- Cloverleaf secondary structure with three hairpin loops and an acceptor stem
- The anticodon loop (at the bottom of the cloverleaf) contains the three-nucleotide anticodon that base-pairs with the mRNA codon
- The acceptor stem (at the 3' end) is where the amino acid attaches, always at a conserved CCA sequence
There are roughly 20 families of tRNA molecules in a cell, one for each standard amino acid. However, most organisms have more than 20 individual tRNA species because of wobble pairing and codon redundancy.
tRNAs are also among the most heavily modified RNA molecules. Post-transcriptional modifications (like pseudouridine and inosine) help stabilize their structure and fine-tune codon recognition.

Ribosomal RNA (rRNA) in Protein Synthesis Machinery
rRNA is the most abundant type of RNA in the cell and forms both the structural scaffold and the catalytic core of the ribosome. It makes up about 60% of the ribosome's total mass, with proteins accounting for the rest.
The rRNA composition differs between eukaryotes and prokaryotes:
| Eukaryotes | Prokaryotes | |
|---|---|---|
| Large subunit rRNAs | 28S, 5.8S, 5S | 23S, 5S |
| Small subunit rRNA | 18S | 16S |
A critical point: rRNA is not just structural. The peptidyl transferase activity that catalyzes peptide bond formation during translation is carried out by the 23S rRNA (in prokaryotes) or 28S rRNA (in eukaryotes), not by a protein enzyme. This makes the ribosome a ribozyme, an RNA molecule with catalytic function.
Because rRNA sequences are highly conserved across species, they're widely used in evolutionary biology and taxonomy. The 16S rRNA gene, for instance, is the standard marker for identifying and classifying bacteria.
Regulatory RNAs
Small Nuclear RNA (snRNA) in Pre-mRNA Processing
snRNAs are essential components of the splicing machinery in eukaryotic cells. They don't work alone; each snRNA associates with specific proteins to form a small nuclear ribonucleoprotein (snRNP), pronounced "snurp."
Multiple snRNPs assemble together on the pre-mRNA to form the spliceosome, the large complex that catalyzes intron removal. The major spliceosomal snRNAs and their roles:
- U1 snRNA recognizes and binds the 5' splice site
- U2 snRNA binds the branch point sequence within the intron
- U4, U5, and U6 snRNAs join to complete the spliceosome and carry out the catalytic steps
A separate minor spliceosome (containing U11, U12, U4atac, and U6atac) processes a small subset of introns with non-canonical splice site sequences. These represent less than 1% of all introns but are found in functionally important genes.
MicroRNA (miRNA) and Gene Expression Regulation
MicroRNAs (miRNAs) are short non-coding RNAs, typically 20-24 nucleotides long, that regulate gene expression after transcription. They act as fine-tuners, dialing down the output of specific genes.
How miRNAs are produced and function:
- A longer precursor RNA (pri-miRNA) is transcribed and forms a hairpin structure
- The enzyme Drosha trims it in the nucleus to a shorter precursor (pre-miRNA)
- After export to the cytoplasm, Dicer cleaves it into the mature ~22-nucleotide miRNA duplex
- One strand is loaded into the RNA-induced silencing complex (RISC)
- The miRNA guides RISC to complementary sequences in the 3' UTR of target mRNAs
The result is either translational repression (the ribosome is blocked) or mRNA degradation, depending on the degree of complementarity. In animals, miRNAs usually bind with imperfect complementarity, leading to translational repression. A single miRNA can regulate hundreds of different mRNA targets, making miRNAs powerful regulators of development, cell differentiation, and disease processes including cancer.
Small Interfering RNA (siRNA) in Gene Silencing
siRNAs are double-stranded RNA molecules, generally 20-25 base pairs long, that silence gene expression through the RNA interference (RNAi) pathway.
The siRNA pathway works as follows:
- Long double-stranded RNA (from viral replication, transposons, or experimental introduction) is cleaved by Dicer into short siRNA duplexes
- The siRNA duplex is loaded into RISC, and one strand (the passenger strand) is discarded
- The remaining guide strand directs RISC to mRNAs with perfectly complementary sequences
- RISC cleaves the target mRNA, leading to its degradation
siRNA vs. miRNA: Both use RISC and Dicer, but they differ in origin and mechanism. siRNAs come from long double-stranded RNA precursors and require near-perfect complementarity to their targets, leading to direct mRNA cleavage. miRNAs come from single-stranded hairpin precursors (processed by Drosha first) and typically bind with imperfect complementarity, causing translational repression.
siRNAs have become a major tool in molecular biology research. By introducing synthetic siRNAs into cells, researchers can selectively knock down specific genes to study their function. Therapeutic siRNAs are also being developed to treat viral infections and genetic disorders, with the first siRNA drug (patisiran) approved in 2018.