Supervised learning is a cornerstone of machine learning in bioinformatics. It uses labeled data to train models that can predict or classify new information, playing a vital role in tasks from genomics to proteomics.
This topic covers the fundamentals, algorithms, and applications of supervised learning in biological contexts. It addresses challenges like high-dimensional data, feature selection, and model evaluation, while also considering ethical implications and interpretability in bioinformatics research.
Fundamentals of supervised learning
- Supervised learning forms a crucial component of machine learning in bioinformatics applications
- Involves training models on labeled data to make predictions or classifications on new, unseen data
- Plays a significant role in various biological data analysis tasks, from genomics to proteomics
Definition and core concepts
- Learning algorithm trained on input-output pairs to predict outputs for new inputs
- Utilizes labeled training data to learn patterns and relationships
- Aims to minimize the difference between predicted and actual outputs
- Involves key concepts like loss functions, optimization algorithms, and model parameters
Types of supervised learning
- Classification predicts discrete class labels or categories
- Regression estimates continuous numerical values
- Ordinal regression predicts ordered categories
- Multi-label classification assigns multiple labels to each instance
- Sequence-to-sequence learning maps input sequences to output sequences
Training vs testing data
- Training data used to teach the model patterns and relationships
- Testing data evaluates model performance on unseen examples
- Validation set helps tune hyperparameters and prevent overfitting
- Data splitting techniques (holdout, k-fold cross-validation) ensure robust evaluation
- Stratified sampling maintains class distribution across splits
Common supervised algorithms
- Supervised learning algorithms form the backbone of many bioinformatics applications
- Different algorithms excel at various tasks and data types in biological research
- Understanding algorithm strengths and weaknesses crucial for effective model selection
Decision trees
- Hierarchical structure of nodes representing features and branches representing decisions
- Splits data based on feature values to create homogeneous subsets
- Advantages include interpretability and handling of both numerical and categorical data
- Prone to overfitting, especially with deep trees
- Widely used in gene expression analysis and protein function prediction
Random forests
- Ensemble method combining multiple decision trees to improve predictive performance
- Each tree trained on a random subset of features and data samples (bagging)
- Reduces overfitting and improves generalization compared to single decision trees
- Effective for high-dimensional biological data (gene expression, genomic sequences)
- Provides feature importance rankings useful for biomarker discovery
Support vector machines
- Finds optimal hyperplane to separate classes in high-dimensional space
- Kernel trick allows non-linear classification by transforming input space
- Effective for protein structure prediction and genomic sequence classification
- Handles high-dimensional data well, making it suitable for many bioinformatics tasks
- Sensitive to feature scaling and choice of kernel function
Neural networks
- Interconnected layers of artificial neurons process and transform input data
- Deep learning architectures with multiple hidden layers capture complex patterns
- Convolutional neural networks excel at sequence and image-based biological data
- Recurrent neural networks handle time-series and sequential data (RNA sequences)
- Requires large amounts of training data and careful hyperparameter tuning
Feature selection and engineering
- Critical process in bioinformatics to handle high-dimensional biological data
- Improves model performance, reduces overfitting, and enhances interpretability
- Crucial for identifying relevant biomarkers and understanding biological mechanisms
Importance of feature selection
- Reduces curse of dimensionality in high-throughput biological data
- Improves model performance by focusing on most informative features
- Enhances interpretability by identifying key biological factors
- Reduces computational complexity and storage requirements
- Helps mitigate overfitting, especially with limited sample sizes
Feature extraction techniques
- Principal Component Analysis (PCA) reduces dimensionality while preserving variance
- Independent Component Analysis (ICA) separates mixed signals in biological data
- Non-negative Matrix Factorization (NMF) useful for gene expression data analysis
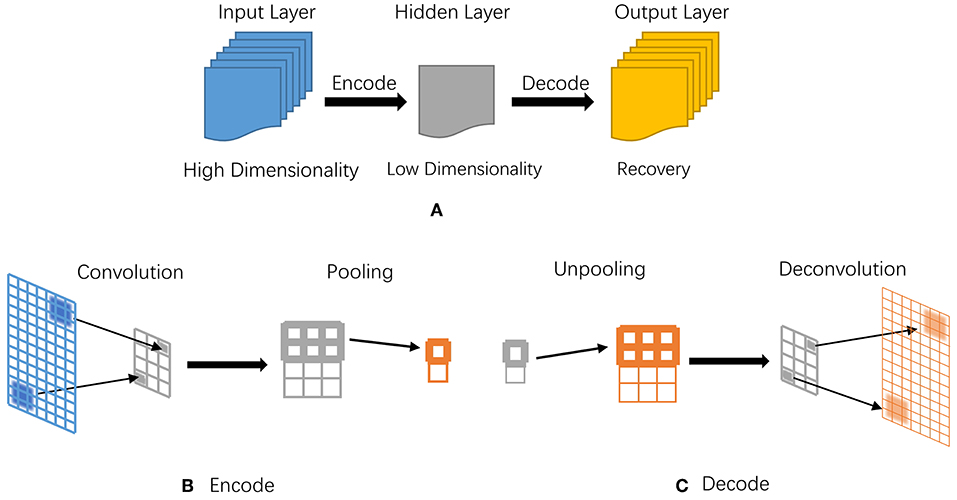
- Autoencoder neural networks learn compact representations of input data
- Domain-specific techniques (protein sequence encoding, structural descriptors)
Dimensionality reduction methods
- t-SNE visualizes high-dimensional data in 2D or 3D space
- UMAP preserves both local and global structure in dimensionality reduction
- Linear Discriminant Analysis (LDA) maximizes class separability
- Feature agglomeration combines similar features based on correlation or distance
- Manifold learning techniques (Isomap, Locally Linear Embedding) for non-linear reduction
Model evaluation and validation
- Crucial for assessing model performance and generalization in bioinformatics
- Ensures reliable and reproducible results in biological data analysis
- Helps identify and mitigate issues like overfitting and dataset bias
Cross-validation techniques
- K-fold cross-validation partitions data into k subsets for multiple train-test cycles
- Leave-one-out cross-validation uses a single sample for testing in each iteration
- Stratified cross-validation maintains class distribution in each fold
- Nested cross-validation for hyperparameter tuning and unbiased performance estimation
- Time series cross-validation for temporally dependent biological data
Performance metrics
- Accuracy measures overall correct predictions but can be misleading with imbalanced data
- Precision, recall, and F1-score provide detailed insights into class-specific performance
- Area Under the ROC Curve (AUC-ROC) evaluates binary classification across thresholds
- Mean Squared Error (MSE) and R-squared assess regression model performance
- Domain-specific metrics (Q3 accuracy for protein secondary structure prediction)
Overfitting vs underfitting
- Overfitting occurs when model learns noise in training data, leading to poor generalization
- Underfitting happens when model fails to capture underlying patterns in the data
- Bias-variance tradeoff balances model complexity and generalization ability
- Regularization techniques (L1, L2) help prevent overfitting
- Learning curves visualize model performance across different training set sizes
Supervised learning in bioinformatics
- Supervised learning techniques play a crucial role in various bioinformatics applications
- Enable prediction and classification tasks across different biological data types
- Contribute to advancing our understanding of complex biological systems and processes
Gene expression analysis
- Differential expression analysis identifies genes with significant changes between conditions
- Gene set enrichment analysis reveals biological pathways associated with gene lists
- Supervised classification of cancer subtypes based on gene expression profiles
- Prediction of gene regulatory networks from time-series expression data
- Identification of biomarkers for disease diagnosis and prognosis
Protein structure prediction
- Secondary structure prediction classifies amino acid residues into structural elements
- Tertiary structure prediction estimates 3D coordinates of protein atoms
- Contact map prediction identifies residue pairs in close spatial proximity
- Protein-protein interaction site prediction locates potential binding regions
- Enzyme function prediction based on sequence and structural features
Genomic sequence classification
- Identification of coding regions (exons) and non-coding regions (introns) in DNA sequences
- Prediction of transcription factor binding sites and regulatory elements
- Classification of genomic variants (SNPs, indels) as pathogenic or benign
- Metagenomic sequence classification for microbial community analysis
- Prediction of splice sites and alternative splicing events in pre-mRNA sequences
Challenges in biological data
- Biological data presents unique challenges for supervised learning applications
- Addressing these challenges crucial for developing robust and reliable models
- Requires specialized techniques and careful consideration of data characteristics
High-dimensionality issues
- Curse of dimensionality affects model performance and interpretability
- Feature selection and dimensionality reduction techniques mitigate high-dimensionality
- Specialized algorithms (Random Forests, SVMs) handle high-dimensional data effectively
- Regularization methods prevent overfitting in high-dimensional spaces
- Ensemble methods combine multiple models to improve performance on high-dimensional data
Imbalanced datasets
- Common in biological data (rare diseases, minority cell types)
- Leads to biased models favoring majority classes
- Resampling techniques (oversampling, undersampling) balance class distributions
- Synthetic data generation (SMOTE) creates new minority class samples
- Cost-sensitive learning assigns higher penalties to misclassifying minority classes
Noise and variability
- Biological data often contains experimental noise and natural variability
- Batch effects introduce systematic biases across experiments or platforms
- Data normalization techniques reduce technical variability
- Robust statistical methods handle outliers and non-normal distributions
- Ensemble methods and data augmentation improve model robustness to noise
Ensemble methods
- Combine multiple models to improve overall performance and robustness
- Particularly effective for complex biological data with high variability
- Reduce overfitting and enhance generalization in bioinformatics applications
Bagging vs boosting
- Bagging (Bootstrap Aggregating) trains models on random subsets of data
- Reduces variance and helps prevent overfitting
- Random Forests use bagging with decision trees
- Boosting trains models sequentially, focusing on misclassified samples
- Gradient Boosting and AdaBoost popular boosting algorithms in bioinformatics
Stacking and blending
- Stacking combines predictions from multiple models using a meta-learner
- Different levels of stacking can capture complex patterns in biological data
- Blending uses a hold-out set to train the meta-learner
- Effective for integrating diverse data types in multi-omics studies
- Can incorporate domain knowledge through carefully designed base models
Voting classifiers
- Combine predictions from multiple models through voting mechanisms
- Hard voting uses majority rule for final classification
- Soft voting weighs model predictions based on their confidence scores
- Weighted voting assigns importance to different models or data sources
- Useful for integrating predictions from different algorithms or data modalities
Hyperparameter tuning
- Critical process for optimizing model performance in bioinformatics applications
- Balances model complexity and generalization ability
- Helps adapt generic algorithms to specific biological data characteristics
Grid search vs random search
- Grid search exhaustively evaluates all combinations of predefined hyperparameter values
- Computationally expensive but guarantees finding the best combination within the grid
- Random search samples hyperparameter values from predefined distributions
- More efficient than grid search, especially for high-dimensional hyperparameter spaces
- Effective for identifying important hyperparameters in biological model optimization
Bayesian optimization
- Sequential model-based optimization technique for efficient hyperparameter tuning
- Uses probabilistic model (Gaussian Process) to guide search towards promising regions
- Balances exploration of unknown areas and exploitation of known good regions
- Particularly useful for computationally expensive bioinformatics models
- Incorporates prior knowledge about hyperparameter importance and ranges
Automated machine learning
- Automates the entire machine learning pipeline, including hyperparameter tuning
- Techniques like Auto-SKLearn and TPOT optimize model selection and hyperparameters
- Neural Architecture Search (NAS) automates design of neural network architectures
- Enables non-experts to apply machine learning to complex biological problems
- Helps standardize and reproduce machine learning workflows in bioinformatics research
Interpretability and explainability
- Crucial for understanding and validating machine learning models in bioinformatics
- Enables biological insights and hypothesis generation from complex models
- Addresses the "black box" nature of some advanced algorithms (deep learning)
Feature importance analysis
- Identifies most influential features in model predictions
- Random Forest feature importance based on decrease in impurity or permutation
- Gradient Boosting feature importance derived from split gain or frequency
- Linear model coefficients indicate feature relevance and direction of influence
- Useful for biomarker discovery and understanding key factors in biological processes
SHAP values
- SHapley Additive exPlanations provide unified approach to feature importance
- Assigns credit to each feature based on game theory principles
- Local and global explanations of model predictions
- Handles interactions between features and non-linear relationships
- Particularly useful for complex biological models with many interacting factors
Model-agnostic interpretation methods
- LIME (Local Interpretable Model-agnostic Explanations) explains individual predictions
- Partial Dependence Plots show average effect of features on model output
- Individual Conditional Expectation plots reveal feature effects for specific instances
- Accumulated Local Effects plots handle correlated features in biological data
- Global Surrogate Models approximate complex models with interpretable ones
Ethical considerations
- Increasing importance as machine learning becomes more prevalent in bioinformatics
- Addresses potential societal impacts and risks of AI in biological and medical research
- Ensures responsible development and application of supervised learning in life sciences
Bias in biological datasets
- Selection bias in data collection can lead to skewed model predictions
- Demographic biases may result in models that perform poorly for underrepresented groups
- Historical biases in medical data can perpetuate existing health disparities
- Careful data collection and preprocessing to mitigate biases
- Regular audits of model performance across different demographic groups
Privacy concerns
- Sensitive nature of genetic and medical data requires robust privacy protections
- De-identification techniques to protect individual privacy in large-scale genomic studies
- Federated learning allows model training without sharing raw data
- Differential privacy adds controlled noise to prevent individual identification
- Secure multi-party computation for collaborative research while preserving data privacy
Reproducibility challenges
- Ensuring reproducibility of machine learning results in bioinformatics research
- Version control for data, code, and model artifacts
- Detailed documentation of data preprocessing, model architecture, and hyperparameters
- Use of standardized benchmarks and evaluation metrics
- Open-source sharing of code and models to facilitate peer review and validation