Non-coding RNAs are crucial players in biology, regulating genes without becoming proteins. They come in various types, including long non-coding RNAs, small RNAs like miRNAs, and circular RNAs, each with unique functions and structures.
Bioinformatics tools are essential for identifying, classifying, and analyzing ncRNAs. These tools use sequence-based, structure-based, and machine learning approaches to predict ncRNAs and their functions, aiding in understanding their roles in gene regulation and disease.
Types of non-coding RNA
- Non-coding RNAs play crucial roles in various biological processes without being translated into proteins
- Bioinformatics approaches enable identification, classification, and functional analysis of ncRNAs
- Understanding ncRNA types aids in developing targeted strategies for gene regulation and disease treatment
Long non-coding RNAs
- Transcripts longer than 200 nucleotides with diverse regulatory functions
- Involved in chromatin remodeling, transcriptional regulation, and post-transcriptional processing
- Includes well-known examples (Xist, HOTAIR, MALAT1)
- Often exhibit tissue-specific expression patterns
- Can act as scaffolds for protein complexes or decoys for other regulatory molecules
Small non-coding RNAs
- Short RNA molecules typically less than 200 nucleotides in length
- Includes microRNAs (miRNAs), small interfering RNAs (siRNAs), and Piwi-interacting RNAs (piRNAs)
- Function in gene silencing through complementary base pairing with target mRNAs
- miRNAs regulate gene expression post-transcriptionally
- siRNAs defend against viral infections and regulate transposable elements
- piRNAs maintain genome stability in germ cells
Circular RNAs
- Covalently closed RNA molecules formed by back-splicing events
- Highly stable due to resistance to exonuclease degradation
- Act as miRNA sponges, regulating miRNA activity
- Involved in protein sequestration and transcriptional regulation
- Some circular RNAs can be translated into proteins
- Emerging roles in various biological processes and diseases
Biological functions of ncRNAs
- ncRNAs participate in diverse cellular processes, impacting gene expression and cellular homeostasis
- Bioinformatics tools help predict and analyze ncRNA functions based on sequence, structure, and interactions
- Understanding ncRNA functions is crucial for developing RNA-based therapeutics and diagnostic tools
Gene regulation mechanisms
- Transcriptional regulation through interaction with promoter regions
- Post-transcriptional regulation via mRNA stability and translation control
- Epigenetic regulation by guiding chromatin modifiers to specific genomic loci
- Competitive endogenous RNA (ceRNA) networks involving miRNA sequestration
- Allosteric regulation of protein function through direct RNA-protein interactions
Epigenetic modifications
- Recruitment of histone-modifying enzymes to specific genomic regions
- DNA methylation patterns influenced by long non-coding RNAs
- X chromosome inactivation mediated by Xist lncRNA
- Imprinting control regions regulated by ncRNAs
- Chromatin remodeling facilitated by ncRNA-protein complexes
Cellular processes involvement
- Cell cycle regulation and apoptosis control
- Differentiation and development of various tissues and organs
- Stress response and cellular homeostasis maintenance
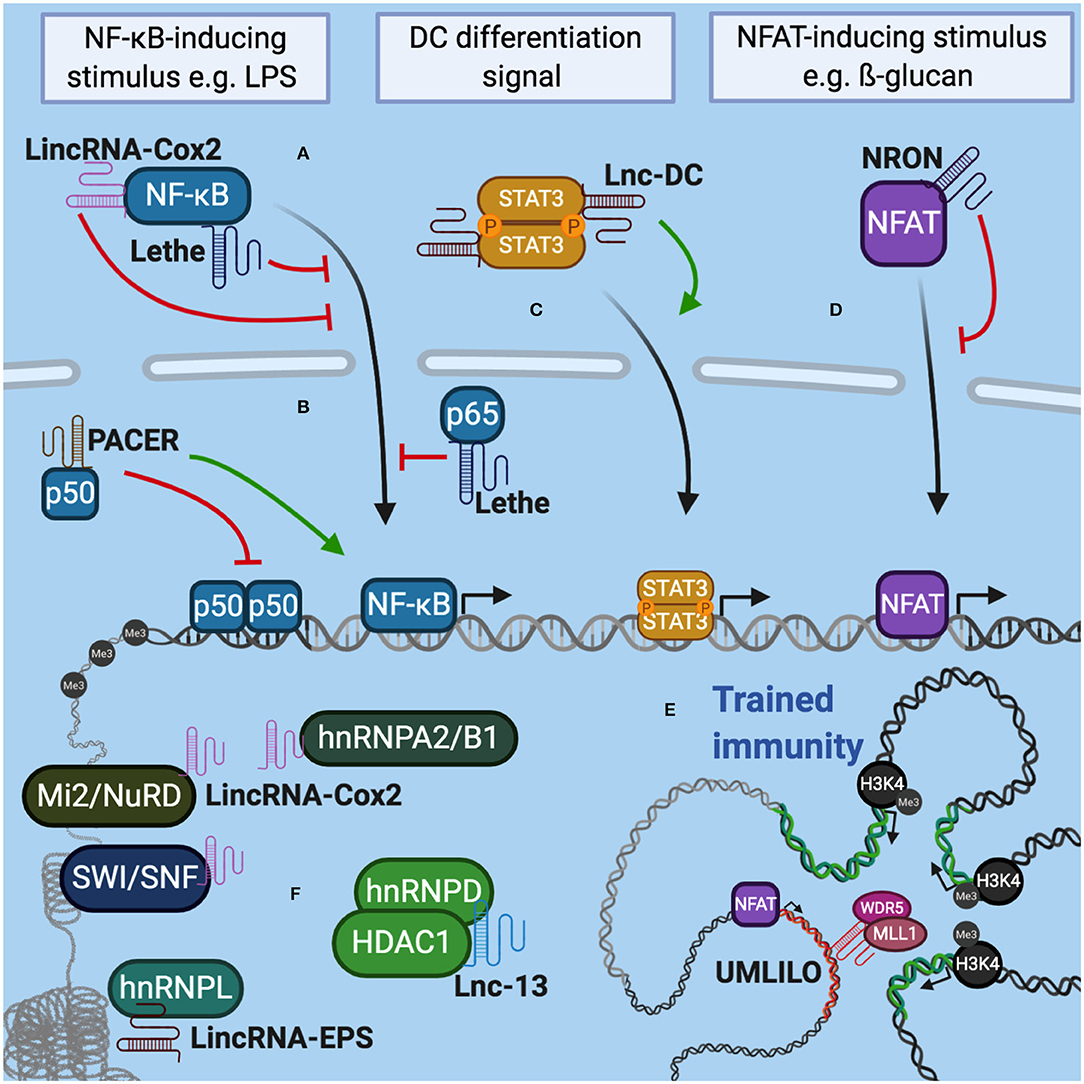
- Immune system modulation and inflammatory processes
- Stem cell pluripotency and lineage commitment
Computational identification methods
- Bioinformatics approaches enable large-scale discovery and annotation of ncRNAs
- Integration of multiple prediction methods improves accuracy in ncRNA identification
- Continuous development of algorithms enhances sensitivity and specificity of ncRNA detection
Sequence-based prediction
- Homology-based methods using BLAST or HMMER for known ncRNA families
- De novo prediction using sequence composition features (GC content, k-mer frequencies)
- Comparative genomics approaches to identify conserved non-coding elements
- Codon substitution frequency (CSF) analysis to distinguish coding from non-coding sequences
- Machine learning classifiers trained on sequence-derived features
Structure-based prediction
- Secondary structure prediction using energy minimization algorithms (RNAfold, Mfold)
- Covariance models to capture both sequence and structure conservation
- Structural motif identification using graph-based algorithms
- Minimum free energy (MFE) calculations to assess RNA stability
- Comparative structure prediction across multiple species
Machine learning approaches
- Support Vector Machines (SVMs) for binary classification of coding vs. non-coding RNAs
- Random Forests for multi-class ncRNA classification
- Deep learning models (Convolutional Neural Networks, Recurrent Neural Networks) for feature extraction
- Ensemble methods combining multiple classifiers for improved accuracy
- Transfer learning to leverage knowledge from well-characterized ncRNAs to predict novel ones
Databases for ncRNA research
- Centralized repositories facilitate access to ncRNA sequences, annotations, and functional information
- Integration of multiple data sources enhances comprehensive analysis of ncRNAs
- Regular updates and curation ensure up-to-date information for researchers
RNA-specific databases
- Rfam database for RNA families and their annotations
- miRBase for microRNA sequences and target predictions
- lncRNAdb for functionally characterized long non-coding RNAs
- circBase for circular RNA sequences and expression data
- NONCODE for comprehensive ncRNA annotation across multiple species
Integrated genomic databases
- UCSC Genome Browser for visualizing ncRNAs in genomic context
- Ensembl for gene annotation including ncRNAs
- NCBI Gene for comprehensive gene information including ncRNAs
- GENCODE for high-quality human and mouse gene annotation
- RNAcentral as a unified resource for all ncRNA types
Species-specific resources
- ENCODE project data for human and model organisms
- modENCODE for Drosophila and C. elegans ncRNA annotations
- PlantRNA database for plant-specific ncRNAs
- FlyBase for Drosophila-specific ncRNA information
- WormBase for C. elegans ncRNA data and functional annotations
Experimental validation techniques
- Experimental validation complements computational predictions of ncRNAs
- Combination of high-throughput and targeted approaches ensures comprehensive characterization
- Integration of experimental data with bioinformatics analysis improves functional annotation
RNA sequencing methods
- RNA-seq for transcriptome-wide profiling of ncRNA expression
- Small RNA-seq for specific detection of miRNAs and other small ncRNAs
- Cap analysis gene expression (CAGE) for precise mapping of transcription start sites
- Poly(A)-independent sequencing methods for non-polyadenylated ncRNAs
- Single-cell RNA-seq for cell-type-specific ncRNA expression analysis
Northern blot analysis
- Size-based separation of RNA molecules on agarose or polyacrylamide gels
- Transfer of RNA to a membrane for hybridization with labeled probes
- Detection of specific ncRNAs using radioactive or non-radioactive probes
- Quantification of relative abundance across different samples or conditions
- Validation of predicted ncRNA transcripts and their sizes
qPCR for ncRNA detection
- Reverse transcription of RNA to cDNA for amplification
- Design of specific primers for ncRNA detection
- Real-time monitoring of amplification using fluorescent dyes or probes
- Relative quantification using reference genes for normalization
- Absolute quantification using standard curves for copy number estimation
Bioinformatics tools for ncRNA
- Specialized software facilitates various aspects of ncRNA analysis
- Integration of multiple tools enables comprehensive characterization of ncRNAs
- Continuous development of algorithms improves accuracy and efficiency in ncRNA research
Sequence alignment tools
- BLAST for homology-based searches of ncRNA sequences
- Clustal Omega for multiple sequence alignment of ncRNAs
- MUSCLE for fast and accurate alignment of large datasets
- T-Coffee for combining global and local alignment methods
- MAFFT for alignment of sequences with long insertions and deletions
Secondary structure prediction
- RNAfold for predicting minimum free energy structures
- Mfold for generating multiple suboptimal structures
- RNAstructure for incorporating experimental constraints in structure prediction
- SHAPE-MaP for high-throughput RNA structure probing and analysis
- RNAalifold for consensus structure prediction from multiple alignments
Expression analysis software
- DESeq2 for differential expression analysis of RNA-seq data
- edgeR for analyzing count-based expression data
- Cufflinks for transcript assembly and quantification
- Salmon for fast and accurate transcript quantification
- sleuth for differential analysis of kallisto pseudoalignments
Evolutionary conservation of ncRNAs
- Evolutionary conservation provides insights into functional importance of ncRNAs
- Comparative genomics approaches reveal conserved ncRNA elements across species
- Integration of conservation data with functional annotations enhances understanding of ncRNA roles
Comparative genomics approaches
- Whole-genome alignments to identify conserved non-coding elements
- Synteny analysis to detect positional conservation of ncRNAs
- Identification of ultraconserved elements in non-coding regions
- Conservation of secondary structure elements across species
- Detection of compensatory mutations maintaining RNA structure
Phylogenetic analysis methods
- Maximum likelihood methods for reconstructing ncRNA evolutionary history
- Bayesian inference for estimating posterior probabilities of phylogenetic trees
- Neighbor-joining algorithms for rapid tree construction
- Parsimony-based methods for inferring ancestral sequences
- Molecular clock analyses to estimate divergence times of ncRNA families
Functional conservation patterns
- Identification of conserved regulatory motifs in ncRNA sequences
- Analysis of conserved RNA-protein interaction sites
- Detection of conserved miRNA target sites across species
- Evolutionary rates of different ncRNA classes and families
- Lineage-specific expansions or losses of ncRNA families
ncRNA interactions
- ncRNAs form complex interaction networks with various biomolecules
- Understanding these interactions is crucial for elucidating ncRNA functions
- Bioinformatics tools aid in predicting and analyzing ncRNA interaction partners
RNA-protein interactions
- CLIP-seq methods for genome-wide mapping of RNA-protein binding sites
- RIP-seq for identifying RNAs associated with specific proteins
- Computational prediction of RNA-binding protein motifs
- Structural analysis of RNA-protein complexes using X-ray crystallography and cryo-EM
- Integration of interaction data with functional annotations to infer biological roles
RNA-DNA interactions
- Triple helix formation between lncRNAs and genomic DNA
- R-loop structures formed by RNA-DNA hybrids
- CHART and ChIRP techniques for mapping RNA-chromatin interactions
- Computational prediction of RNA-DNA interaction potential
- Functional consequences of RNA-DNA interactions on gene expression and chromatin structure

RNA-RNA interactions
- Base-pairing interactions between miRNAs and target mRNAs
- Long-range interactions in RNA secondary structures
- Competitive endogenous RNA networks involving multiple RNA species
- CLASH and PARIS methods for high-throughput RNA-RNA interaction mapping
- Computational tools for predicting RNA-RNA interactions (IntaRNA, RNAup)
Functional annotation of ncRNAs
- Assigning functions to ncRNAs is crucial for understanding their biological roles
- Integration of multiple data types improves accuracy of functional predictions
- Bioinformatics approaches enable large-scale functional annotation of ncRNAs
Gene ontology enrichment
- Analysis of GO terms associated with genes co-expressed with ncRNAs
- Functional enrichment of predicted ncRNA targets
- Development of RNA-specific GO terms and annotations
- Integration of GO enrichment results with expression data
- Visualization tools for exploring GO enrichment results
Pathway analysis
- KEGG pathway mapping of genes associated with ncRNAs
- Reactome pathway analysis for detailed biological processes
- Identification of signaling pathways regulated by ncRNAs
- Integration of pathway information with ncRNA expression data
- Network-based pathway analysis incorporating ncRNA interactions
Network-based approaches
- Construction of ncRNA-mRNA co-expression networks
- Protein-protein interaction networks incorporating ncRNA data
- Regulatory networks integrating transcription factors and ncRNAs
- Identification of network motifs involving ncRNAs
- Centrality measures to identify key ncRNAs in biological networks
Disease associations of ncRNAs
- ncRNAs play crucial roles in various diseases, offering potential as biomarkers and therapeutic targets
- Bioinformatics approaches aid in identifying disease-associated ncRNAs and their mechanisms
- Integration of clinical data with ncRNA profiles enhances understanding of disease processes
Cancer-related ncRNAs
- Oncogenic and tumor suppressor roles of lncRNAs in cancer progression
- miRNA dysregulation in various cancer types
- Circular RNAs as potential cancer biomarkers
- ncRNA involvement in metastasis and drug resistance
- Pan-cancer analysis of ncRNA expression patterns
Neurological disorders
- lncRNAs in neurodegenerative diseases (Alzheimer's, Parkinson's)
- miRNA regulation of synaptic plasticity and neuronal function
- ncRNA roles in neurodevelopmental disorders (autism, schizophrenia)
- Circular RNAs in brain function and neurological diseases
- Blood-based ncRNA biomarkers for neurological disorders
Cardiovascular diseases
- lncRNAs in cardiac remodeling and heart failure
- miRNA regulation of lipid metabolism and atherosclerosis
- Circular RNAs in vascular function and disease
- ncRNA biomarkers for myocardial infarction and stroke
- Therapeutic potential of ncRNAs in cardiovascular diseases
Therapeutic potential of ncRNAs
- ncRNAs offer promising avenues for developing novel therapeutic strategies
- Bioinformatics tools aid in designing and optimizing ncRNA-based therapeutics
- Integration of computational and experimental approaches enhances therapeutic development
RNA-based therapeutics
- Antisense oligonucleotides for targeting disease-associated ncRNAs
- miRNA mimics and inhibitors for modulating gene expression
- CRISPR-Cas13 systems for targeted RNA degradation
- RNA aptamers as therapeutic agents and delivery vehicles
- Challenges in RNA stability and delivery for therapeutic applications
Gene therapy applications
- Viral vector-mediated delivery of therapeutic ncRNAs
- Non-viral delivery systems (nanoparticles, liposomes) for ncRNA therapeutics
- Ex vivo gene therapy approaches using engineered ncRNAs
- Tissue-specific promoters for controlled expression of therapeutic ncRNAs
- Genome editing of ncRNA loci for long-term therapeutic effects
Diagnostic biomarkers
- Circulating ncRNAs as non-invasive biomarkers for disease detection
- Tissue-specific ncRNA signatures for cancer diagnosis and prognosis
- Machine learning approaches for developing ncRNA-based diagnostic models
- Integration of ncRNA biomarkers with other molecular and clinical data
- Challenges in standardization and clinical validation of ncRNA biomarkers
Challenges in ncRNA research
- Ongoing technological and methodological advancements address current limitations in ncRNA research
- Bioinformatics plays a crucial role in overcoming challenges through improved algorithms and data integration
- Collaborative efforts between experimental and computational researchers drive progress in the field
Experimental validation difficulties
- Low expression levels of many ncRNAs challenging detection
- Tissue-specific and condition-dependent expression patterns
- Functional redundancy among ncRNAs complicating knockout studies
- Technical challenges in manipulating long non-coding RNAs
- Need for high-throughput methods to validate computationally predicted ncRNAs
Computational prediction limitations
- False positives in de novo ncRNA prediction algorithms
- Difficulty in distinguishing functional ncRNAs from transcriptional noise
- Challenges in predicting functions of novel ncRNAs without homology
- Computational resources required for genome-wide ncRNA analyses
- Integration of heterogeneous data types for improved prediction accuracy
Functional characterization hurdles
- Complexity of ncRNA-mediated regulatory networks
- Subtle phenotypes associated with many ncRNA perturbations
- Challenges in determining direct vs. indirect effects of ncRNAs
- Limited understanding of structure-function relationships in ncRNAs
- Need for improved methods to study ncRNA-protein and ncRNA-DNA interactions