Domains

Archaea and Bacteria
Both Archaea and Bacteria are prokaryotes, meaning their cells lack a true nucleus. Despite sharing that prokaryotic cell structure, they are genetically and biochemically distinct enough to occupy separate domains.
Archaea are single-celled microorganisms best known for thriving in extreme environments like hot springs, salt lakes, and deep-sea hydrothermal vents (though many also live in moderate environments like soil and ocean water). Their cell walls and membranes contain unique lipids not found in Bacteria or Eukarya. These unusual membrane lipids are part of what makes them so resilient in harsh conditions.
Bacteria are single-celled microorganisms found in nearly every environment on Earth, from soil and water to the inside of your digestive tract. A key distinguishing feature is that bacterial cell walls contain peptidoglycan, a complex polymer of amino acids and sugars. Archaea lack peptidoglycan entirely, which is one of the structural differences that supports placing them in a separate domain.
Eukarya
Eukarya includes all organisms whose cells contain a true nucleus and membrane-bound organelles. This domain covers enormous diversity:
- Protists (mostly unicellular eukaryotes like amoebas and algae)
- Fungi (yeasts, molds, mushrooms)
- Plants
- Animals
Eukaryotic cells are structurally more complex than prokaryotic cells. Their DNA is enclosed within a nuclear membrane, and specialized organelles like mitochondria, chloroplasts (in plants and algae), the endoplasmic reticulum, and the Golgi apparatus carry out specific cellular functions. A cytoskeleton of protein filaments provides internal structure and enables cell movement.
Cell Types
Prokaryotic Cells
Prokaryotic cells belong to the domains Archaea and Bacteria. Their defining features include:
- No nucleus or membrane-bound organelles. DNA exists as a single circular chromosome located in the cytoplasm, in a region called the nucleoid.
- Smaller ribosomes (70S) compared to eukaryotic ribosomes.
- Reproduction by binary fission, a form of asexual reproduction where the cell copies its DNA and splits into two identical daughter cells.
Eukaryotic Cells
Eukaryotic cells are found in all members of domain Eukarya. They differ from prokaryotic cells in several important ways:
- DNA is enclosed within a nuclear membrane, organized into linear chromosomes (rather than a single circular chromosome).
- Larger ribosomes (80S) compared to prokaryotic ribosomes.
- Membrane-bound organelles compartmentalize cellular functions (mitochondria for energy, chloroplasts for photosynthesis in plants, lysosomes for digestion, etc.).
- Reproduction occurs through mitosis (for growth and asexual reproduction) or meiosis (which produces gametes for sexual reproduction).
A quick comparison to keep straight: Prokaryotes have 70S ribosomes, no nucleus, and reproduce by binary fission. Eukaryotes have 80S ribosomes, a membrane-bound nucleus, and reproduce by mitosis or meiosis.
Classification Methodology
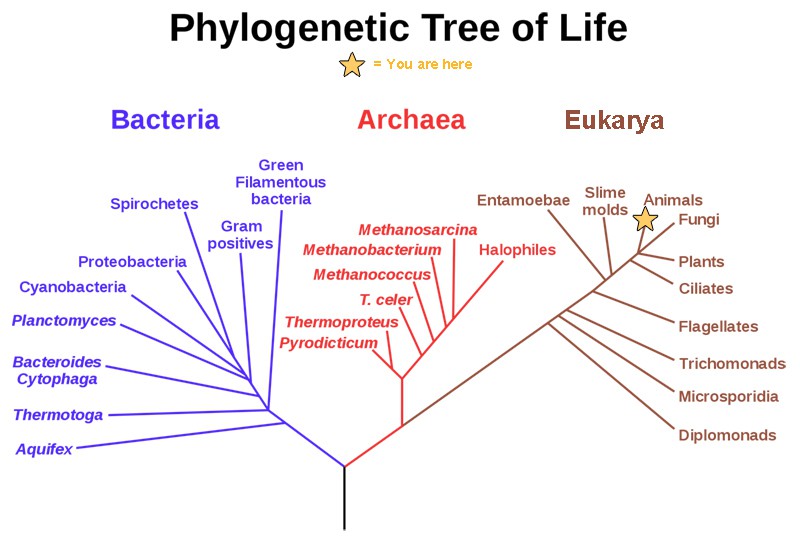
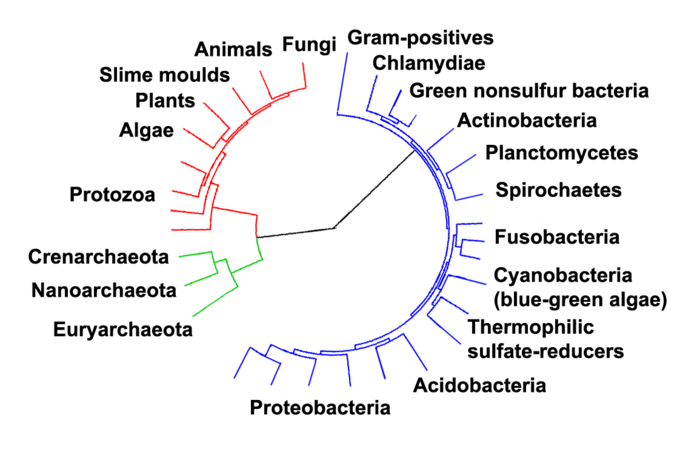
Carl Woese's Contributions
Before Carl Woese's work in the late 1970s, biologists grouped all life into broad kingdoms, and prokaryotes were lumped together as a single group. Woese, an American microbiologist, challenged this by using molecular evidence rather than just physical characteristics to classify organisms.
His key insight was that comparing the sequences of specific genes could reveal deep evolutionary relationships that morphology alone could not. By analyzing ribosomal RNA sequences across hundreds of organisms, he discovered that prokaryotes actually fall into two fundamentally different groups. This led him to propose the three-domain system: Archaea, Bacteria, and Eukarya.
16S rRNA Analysis
The molecule Woese used was 16S rRNA, a component of the small subunit of prokaryotic ribosomes. Here's why it works so well for classification:
- It's universal. Every prokaryote has a gene encoding 16S rRNA (eukaryotes have an analogous molecule, 18S rRNA), so you can compare organisms across all of life.
- It's highly conserved. The gene changes slowly over evolutionary time, meaning even distantly related organisms share recognizable sequence similarities.
- It contains variable regions. While the overall structure is conserved, certain sections accumulate mutations at a steady rate, acting as a molecular clock for measuring evolutionary divergence.
By comparing 16S rRNA sequences, Woese showed that Archaea are as genetically different from Bacteria as both are from Eukarya. That finding is what justified creating three separate domains rather than two.
Horizontal Gene Transfer
Horizontal gene transfer (HGT) is the movement of genetic material between organisms outside of normal parent-to-offspring (vertical) inheritance. It's especially common among prokaryotes and can happen through mechanisms like conjugation, transformation, and transduction.
HGT has real-world consequences. It's a major driver of antibiotic resistance spreading through bacterial populations: one bacterium can acquire a resistance gene from a completely different species.
For classification, HGT creates a genuine challenge. If organisms are swapping genes across species boundaries, their genomes become a mosaic of different evolutionary histories. A phylogenetic tree built from one gene might tell a different story than a tree built from another gene. This is why Woese's choice of 16S rRNA was so important: ribosomal RNA genes are rarely transferred horizontally, making them a more reliable marker for tracing true evolutionary descent.