The Genetic Code
Characteristics of the Genetic Code
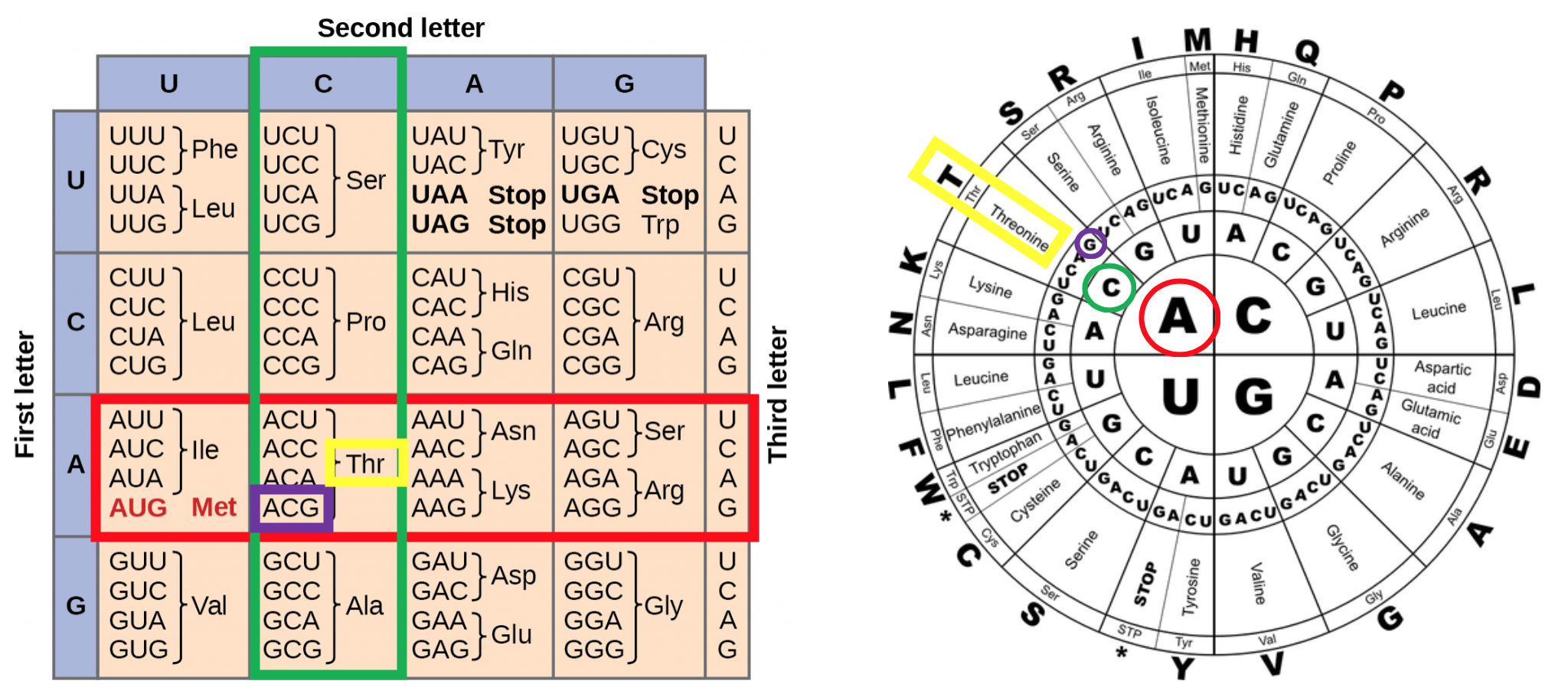
The genetic code is the set of rules cells use to translate nucleotide sequences in mRNA into amino acid sequences in proteins. Every three nucleotides form a codon, and each codon specifies either an amino acid or a signal to start or stop translation.
Here are the key properties you need to know:
- Unambiguous: Each codon codes for exactly one amino acid (or one stop/start signal). There's no confusion about what a given codon means.
- Degenerate: Multiple codons can code for the same amino acid. For example, leucine is specified by six different codons (UUA, UUG, CUU, CUC, CUA, CUG). These are called synonymous codons.
- Comma-less: There's no punctuation or spacer between codons. The ribosome reads the mRNA in a continuous stream of triplets.
- Non-overlapping: Each nucleotide belongs to only one codon. The ribosome moves three nucleotides at a time without reusing any.
- Nearly universal: Almost every organism on Earth uses the same genetic code. A few exceptions exist, most notably in mitochondrial genomes, but the code is remarkably conserved from bacteria to humans.
Three codons serve as stop signals (UAA, UAG, UGA), and one codon (AUG) serves double duty as both the start codon and the codon for methionine. Because the code is read in a fixed frame starting from AUG, shifting the reading frame by even one nucleotide changes every downstream codon and produces a completely different (usually nonfunctional) protein.

Structure and Function of tRNA
Transfer RNA (tRNA) is the adaptor molecule that physically connects an mRNA codon to its corresponding amino acid. Without tRNA, there would be no way to "read" the genetic code and convert it into protein.
Secondary structure (cloverleaf): tRNA folds into a cloverleaf shape with four distinct arms:
- Acceptor arm: The 3' end where the amino acid attaches. All tRNAs end with the sequence CCA at this site.
- Anticodon arm: Contains the three-nucleotide anticodon that base-pairs with the mRNA codon during translation.
- D arm: Named for its dihydrouridine residues. Contributes to the overall 3D folding of the molecule.
- TΨC arm: Named for its ribothymidine-pseudouridine-cytidine sequence. Also helps stabilize the tertiary structure.
Tertiary structure (L-shape): The cloverleaf folds further into an L-shaped 3D structure. The amino acid attachment site sits at one end of the L, and the anticodon loop sits at the other. This shape is critical because it positions the anticodon to interact with mRNA in the ribosome's decoding center while the amino acid end reaches into the peptidyl transferase center where peptide bonds form.
tRNA also contains several modified nucleotides (such as inosine and pseudouridine) that increase its structural stability and fine-tune its interactions during translation.
tRNA and Codon-Anticodon Interactions
Codon-Anticodon Pairing
During translation, the tRNA anticodon base-pairs with the mRNA codon in an antiparallel orientation. For example, the mRNA codon 5'-GGC-3' pairs with the tRNA anticodon 3'-CCG-5'. This complementary pairing is what ensures the correct amino acid gets added to the growing polypeptide chain.
The ribosome checks this pairing at its A site (aminoacyl site). If the codon-anticodon match is correct, the ribosome accepts the tRNA and the amino acid is transferred to the polypeptide. If the match is wrong, the tRNA is rejected. This proofreading step is a major reason translation is so accurate.
Wobble Base Pairing
Wobble pairing refers to relaxed base-pairing rules at the third position of the codon (which pairs with the first position of the anticodon). Francis Crick proposed this concept to explain how the degeneracy of the genetic code actually works at the molecular level.
Standard Watson-Crick pairing is strict (A-U, G-C), but at the wobble position, non-standard pairs are tolerated. For instance, inosine (I) in the first position of the anticodon can pair with U, C, or A in the third position of the codon.
Why this matters:
- A single tRNA can recognize multiple synonymous codons. For example, one valine tRNA can read GUU, GUC, GUA, and GUG.
- This reduces the total number of tRNA species a cell needs. Instead of requiring 61 different tRNAs (one per sense codon), cells typically have around 45.
- Wobble pairing explains why the genetic code is degenerate: most of the variation between synonymous codons occurs at the third position, exactly where wobble permits flexibility.
The pattern to remember: the first two positions of a codon are read strictly, but the third position tolerates wobble. That's why synonymous codons almost always differ only at position three.