Genetics and population dynamics form the backbone of evolutionary biology, connecting individual traits to large-scale changes in populations. Stochastic processes are central here because random events like mutations and genetic drift shape genetic diversity, while environmental fluctuations drive changes in population size. This unit ties those stochastic tools to real biological systems.
Genetics Basics
Genetics is the study of heredity and variation in living organisms. Before you can model how allele frequencies shift stochastically in a population, you need a solid grasp of how genetic information is structured, inherited, and expressed.
DNA Structure and Function
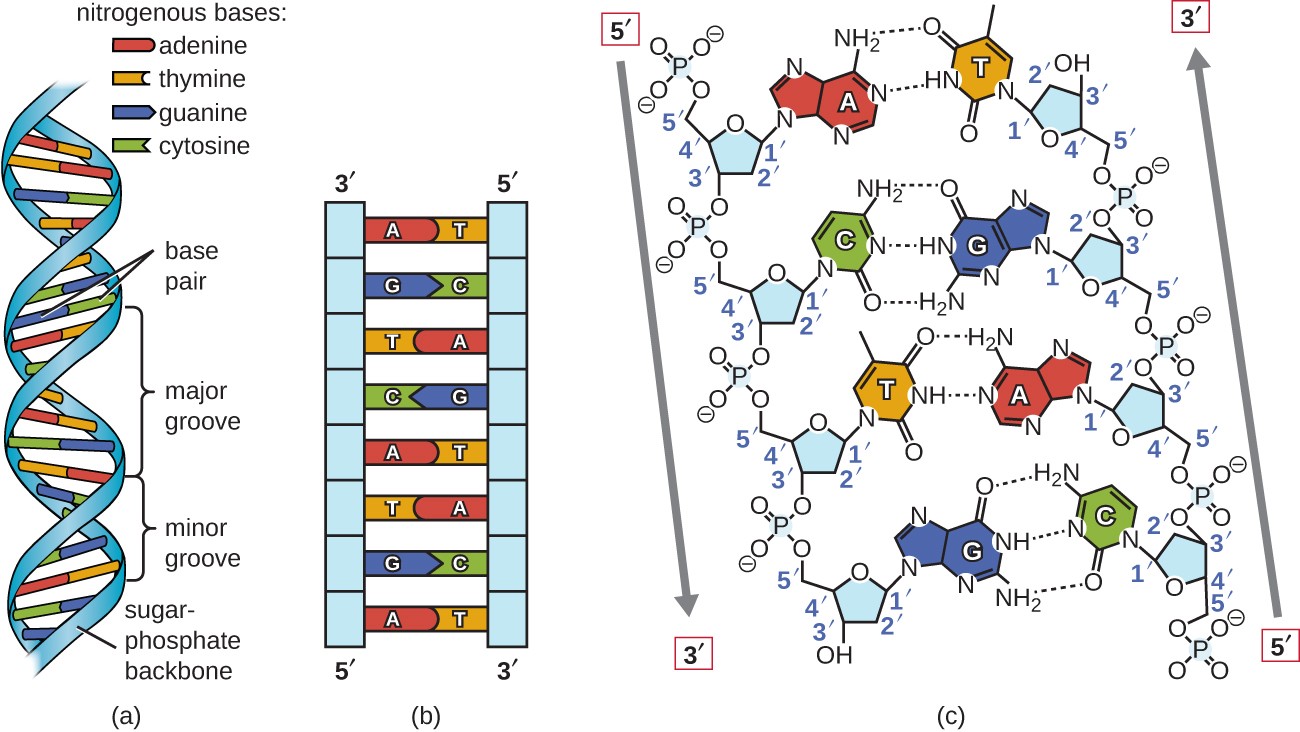
DNA (deoxyribonucleic acid) carries hereditary information in most organisms. It's composed of four nucleotide bases: adenine (A), thymine (T), guanine (G), and cytosine (C). These bases pair specifically (A with T, G with C) to form the rungs of the double helix, while the sugar-phosphate backbone forms the sides.
The sequence of bases along the DNA molecule encodes instructions for synthesizing proteins and regulating cellular functions. This sequence is what stochastic mutation models act upon.
Genes and Alleles
A gene is a segment of DNA that codes for a specific trait or function, located at a particular position (locus) on a chromosome. Alleles are different versions of a gene that can exist at a given locus. For example, a gene controlling flower color might have a "purple" allele and a "white" allele.
An individual inherits one allele from each parent, and the combination of alleles at a locus constitutes that individual's genotype for that gene.
Genotypes and Phenotypes
- The genotype is the specific set of alleles an individual carries at each locus
- The phenotype is the observable trait that results from the interaction between genotype and environment
- Dominant alleles mask the expression of recessive alleles, so the dominant trait appears in the phenotype whenever at least one dominant allele is present
This distinction matters for stochastic modeling because selection acts on phenotypes, but drift and mutation act on the underlying allele frequencies.
Mendelian Inheritance Patterns
Gregor Mendel's experiments with pea plants established the core principles of inheritance:
- Law of segregation: Allele pairs separate during gamete formation, so each gamete receives one allele from each pair
- Law of independent assortment: The inheritance of one gene is independent of the inheritance of another (assuming they're on different chromosomes)
Inheritance patterns include dominant-recessive, codominant, and incomplete dominance. These patterns determine how genotype frequencies translate into phenotype frequencies, which feeds directly into population genetics models.
Population Genetics
Population genetics studies genetic variation within and among populations, focusing on how allele frequencies change over time. This is where stochastic processes become especially important, since random sampling effects can shift allele frequencies even without selection.
Allele Frequencies in Populations
Allele frequency is the proportion of a specific allele relative to all alleles at that locus in a population. If there are two alleles at a locus, their frequencies must sum to 1. For instance, if the frequency of allele A is 0.7, the frequency of allele a is 0.3.
Changes in allele frequencies over time can result from mutation, migration, genetic drift, and natural selection. Tracking these changes is the central task of population genetics.
Hardy-Weinberg Equilibrium
The Hardy-Weinberg principle provides a null model: allele frequencies remain constant across generations when no evolutionary forces are acting. The equilibrium equation is:
Here and are the frequencies of two alleles at a locus. The terms , , and give the expected genotype frequencies for homozygous dominant, heterozygous, and homozygous recessive individuals, respectively.
The assumptions required for Hardy-Weinberg equilibrium are:
- Large (effectively infinite) population size
- Random mating
- No mutation
- No migration
- No natural selection
When any of these assumptions is violated, allele frequencies change. Stochastic models specifically address what happens when assumption 1 fails (finite populations introduce drift) and when assumptions 3-4 introduce random variation.
Factors Affecting Allele Frequencies
- Mutation introduces new alleles, though mutation rates are typically low (on the order of per base pair per generation in humans)
- Migration (gene flow) can introduce new alleles or shift existing frequencies when individuals move between populations
- Non-random mating (assortative mating, inbreeding) alters genotype frequencies by increasing homozygosity, even if allele frequencies themselves don't change immediately
- Natural selection favors alleles that increase fitness, systematically shifting allele frequencies toward adaptive outcomes
Genetic Drift vs. Natural Selection
These two forces are fundamentally different in character:
- Genetic drift is a stochastic process: random sampling of alleles during reproduction causes allele frequencies to fluctuate unpredictably, especially in small populations. Drift can lead to fixation (frequency reaches 1) or loss (frequency reaches 0) purely by chance.
- Natural selection is a deterministic force that systematically favors alleles conferring higher fitness.
In large populations, selection tends to dominate. In small populations, drift can overpower selection, causing even beneficial alleles to be lost or deleterious alleles to become fixed. The relative strength of drift versus selection depends on the product , where is the effective population size and is the selection coefficient. When , drift dominates; when , selection dominates.
Genetic Variation
Genetic variation refers to differences in DNA sequences among individuals within or between populations. It's the raw material that both drift and selection act upon, and without it, populations cannot adapt to changing environments.
Sources of Genetic Variation
- Mutation is the ultimate source, creating new alleles through changes in DNA sequences
- Recombination during meiosis shuffles existing alleles into new combinations on chromosomes
- Sexual reproduction brings together alleles from different parents, increasing the diversity of genotype combinations in offspring
Mutations and Recombination
Mutations come in several forms:
- Point mutations: single nucleotide changes, which can be silent (no amino acid change), missense (different amino acid), or nonsense (premature stop codon)
- Insertions and deletions: addition or removal of nucleotides
- Chromosomal rearrangements: larger-scale structural changes
Recombination occurs during prophase I of meiosis, when homologous chromosomes exchange DNA segments (crossing over). Recombination rates vary along chromosomes, with certain regions acting as hotspots. From a stochastic modeling perspective, recombination breaks up linkage between alleles, which affects how drift and selection act on linked loci.
Quantifying Genetic Variation
- Heterozygosity: the proportion of individuals heterozygous at a given locus, commonly used as a summary measure of variation
- Nucleotide diversity (): the average number of nucleotide differences per site between two randomly chosen sequences in a population
- Measurement techniques include DNA sequencing, restriction fragment length polymorphisms (RFLPs), and microsatellite markers
Implications of Genetic Diversity
High genetic diversity allows populations to adapt to environmental changes and resist disease outbreaks. Low diversity increases the risk of inbreeding depression (reduced fitness from mating between relatives) and raises extinction probability.
Conservation efforts frequently aim to maintain or restore genetic diversity in threatened populations. Stochastic models help quantify how quickly small populations lose diversity through drift, which directly informs management decisions.
Population Dynamics Models
Population dynamics models describe how populations change in size over time, incorporating birth rates, death rates, and environmental constraints. Adding stochastic elements to these models captures the randomness inherent in real biological systems.
Exponential Population Growth
Exponential growth occurs when a population increases at a constant per capita rate, producing a J-shaped curve:
where is population size at time , is the initial population size, is the intrinsic growth rate, and is the base of the natural logarithm.
This model is unrealistic for long time horizons because it assumes unlimited resources. However, it's a useful baseline and can approximate growth in populations that are well below carrying capacity.
Logistic Population Growth
The logistic model adds a carrying capacity constraint, producing an S-shaped curve:
where is the carrying capacity. As approaches , the growth rate slows toward zero. This is a deterministic model, but it serves as the foundation for stochastic extensions.
Carrying Capacity and Limiting Factors
Carrying capacity () is the maximum population size an environment can sustain indefinitely. Limiting factors constrain growth and include food availability, space, water, and other resources.
As populations approach , density-dependent factors like competition, predation, and disease become increasingly important in regulating growth. In stochastic models, itself may fluctuate randomly due to environmental variation.

Stochastic Population Models
Deterministic models give you a single trajectory. Stochastic models give you a distribution of possible outcomes, which is far more realistic. Two key types of randomness are incorporated:
- Demographic stochasticity: random variation in individual birth and death events. Even if the average birth rate is 0.5, any given individual either reproduces or doesn't. This matters most in small populations.
- Environmental stochasticity: random fluctuations in environmental conditions (drought, disease outbreaks) that affect the entire population's growth rate simultaneously.
Stochastic models are used to estimate extinction probabilities and population viability under different management scenarios. For example, a population viability analysis might run thousands of simulated trajectories to determine the probability that a population of 50 individuals persists for 100 years.
Evolutionary Processes
Evolutionary processes describe how populations change over time due to the interplay of genetic variation, selection, drift, and other forces. Understanding these processes is essential for predicting long-term population dynamics.
Fitness and Selection
Fitness measures an individual's relative reproductive success, determined by its genotype and the environment. Natural selection is the differential survival and reproduction of individuals based on fitness differences.
Three major modes of selection shape phenotype distributions:
- Directional selection: favors one extreme phenotype, shifting the population mean over time
- Stabilizing selection: favors intermediate phenotypes, reducing variation
- Disruptive selection: favors both extremes over intermediate phenotypes, potentially creating a bimodal distribution
Types of Natural Selection
- Positive selection increases the frequency of advantageous alleles
- Purifying (negative) selection removes deleterious alleles
- Balancing selection maintains multiple alleles, often through heterozygote advantage (e.g., sickle cell trait conferring malaria resistance) or frequency-dependent selection
- Sexual selection favors traits that increase mating success, even at a survival cost
Adaptation and Speciation
Adaptation is the process by which populations become better suited to their environment through natural selection. Adaptations can be morphological, physiological, or behavioral.
Speciation is the formation of new species through reproductive isolation and genetic divergence:
- Allopatric speciation: populations become geographically separated and diverge over time
- Sympatric speciation: populations diverge without geographic isolation, often through ecological or behavioral differences
Stochastic models of speciation often incorporate both drift and selection to predict how quickly reproductive isolation develops.
Phylogenetics and Evolutionary Trees
Phylogenetics studies evolutionary relationships among species or populations. Phylogenies (evolutionary trees) depict branching patterns of descent from common ancestors, inferred from morphological, behavioral, or molecular data.
Common methods for constructing phylogenetic trees include:
- Maximum parsimony: finds the tree requiring the fewest evolutionary changes
- Maximum likelihood: finds the tree that makes the observed data most probable under a given model of evolution
- Bayesian inference: combines prior information with the data to estimate posterior probabilities of different trees
These methods increasingly rely on stochastic models of sequence evolution to account for the randomness inherent in mutation and substitution processes.
Applications of Population Genetics
Population genetics principles have practical applications across conservation biology, epidemiology, forensics, and agriculture. Stochastic models are frequently used when applying these concepts to real-world problems.
Conservation Genetics
Conservation genetics uses genetic data to manage and protect threatened species. Genetic diversity is assessed to evaluate inbreeding risk and adaptive potential. Genetic rescue involves introducing individuals from a different population to boost diversity and fitness. For example, the Florida panther population was rescued from severe inbreeding depression by introducing Texas pumas in 1995.
Stochastic population models predict the viability of small populations and help determine minimum viable population sizes for conservation planning.
Genetic Epidemiology
Genetic epidemiology studies how genetic factors influence disease distribution in populations. Genome-wide association studies (GWAS) scan the genome for variants associated with complex diseases, and genetic risk scores aggregate many small-effect variants to predict disease susceptibility.
Important methodological considerations include population stratification (confounding due to ancestry differences), cryptic relatedness among study participants, and ascertainment bias in how samples are collected.
Forensics and Paternity Testing
Forensic genetics uses DNA evidence for individual identification and establishing familial relationships:
- Short tandem repeats (STRs) are highly variable markers used for DNA profiling
- Paternity testing compares a child's DNA profile with an alleged father's to calculate the likelihood of paternity
- Population allele frequencies are used to compute the probability of a random match or the paternity index
The statistical calculations underlying forensic genetics rely on population genetics principles, particularly allele frequency estimation.
Crop and Livestock Breeding
Breeding programs aim to improve traits like yield, quality, and disease resistance:
- Marker-assisted selection (MAS) uses genetic markers linked to favorable alleles to guide breeding decisions
- Genomic selection predicts breeding values from an individual's entire genome rather than specific markers, enabling more accurate selection
- Genetic diversity must be managed carefully to balance selection progress against inbreeding depression
Stochastic models help optimize breeding strategies by predicting the long-term response to selection and the rate of diversity loss over multiple generations.