Biological databases are essential tools for storing and organizing vast amounts of biological information. They come in various types, including primary, secondary, sequence, structure, and functional databases, each serving specific research needs in molecular biology and genetics.
These databases play a crucial role in modern biological research, enabling scientists to access, analyze, and interpret complex data. From genomic sequences to protein structures and metabolic pathways, biological databases provide a foundation for advancing our understanding of life at the molecular level.
Types of Biological Databases
Primary and Secondary Databases
- Primary databases contain experimentally derived data directly submitted by researchers
- Include raw sequence data, structural information, and functional annotations
- Examples include GenBank, EMBL, and DDBJ for nucleotide sequences
- Secondary databases compile and analyze information from primary databases
- Provide curated and value-added information
- Often include computational analyses, predictions, and cross-references
- Examples include RefSeq for curated gene sequences and UniProtKB/Swiss-Prot for manually annotated protein information
Sequence and Structure Databases
- Sequence databases store and organize biological sequence information
- Contain DNA, RNA, or protein sequences
- Allow researchers to compare and analyze genetic material across species
- Examples include GenBank (nucleotide sequences) and UniProtKB (protein sequences)
- Structure databases focus on three-dimensional molecular structures
- Store information about the spatial arrangement of atoms in biological molecules
- Crucial for understanding protein function and drug design
- Examples include Protein Data Bank (PDB) for macromolecular structures and Nucleic Acid Database (NaDB) for nucleic acid structures
Functional Databases
- Functional databases provide information on biological roles and interactions
- Contain data on gene expression, protein-protein interactions, and metabolic pathways
- Help researchers understand the complex relationships between biological components
- Examples include Gene Ontology (GO) for gene function annotations and KEGG for metabolic pathways
Specific Database Categories
Genomic Databases
- Store and organize information related to genomes of various organisms
- Include whole genome sequences, gene annotations, and genetic variations
- Facilitate comparative genomics and evolutionary studies
- Examples include Ensembl for vertebrate genomes and FlyBase for Drosophila genomics
- Provide tools for genome browsing, sequence alignment, and variant analysis
- Allow researchers to visualize genomic features and compare across species
- Support identification of disease-associated genes and genetic markers
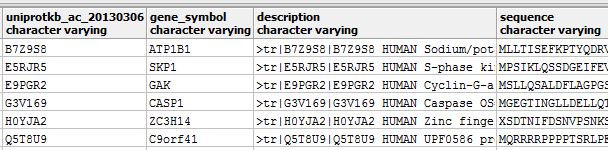
Proteomic Databases
- Focus on protein-related information and analyses
- Store protein sequences, structures, functions, and interactions
- Support research in protein characterization and functional genomics
- Examples include UniProtKB for comprehensive protein information and IntAct for protein interaction data
- Offer tools for protein sequence analysis and structure prediction
- Enable researchers to identify protein domains, motifs, and post-translational modifications
- Facilitate studies on protein evolution and structure-function relationships
Metabolomic Databases
- Contain information on metabolites and metabolic pathways
- Store data on small molecules involved in cellular processes
- Support research in metabolomics and systems biology
- Examples include HMDB (Human Metabolome Database) for human metabolites and MetaCyc for metabolic pathways
- Provide tools for metabolite identification and pathway analysis
- Allow researchers to explore biochemical reactions and metabolic networks
- Facilitate studies on cellular metabolism and metabolic disorders
Major Database Resources

National Center for Biotechnology Information (NCBI)
- Comprehensive resource for molecular biology and genetics information
- Hosts numerous databases covering various aspects of biological research
- Provides tools for data analysis, visualization, and retrieval
- Key databases within NCBI include:
- GenBank for nucleotide sequences
- PubMed for biomedical literature
- BLAST for sequence similarity searches
- Offers resources for researchers, clinicians, and educators
- Supports genomic research, drug discovery, and personalized medicine
- Provides educational materials and training resources
Universal Protein Resource (UniProt)
- Central repository for protein sequence and functional information
- Combines data from Swiss-Prot, TrEMBL, and PIR databases
- Provides comprehensive and non-redundant protein information
- Features of UniProt include:
- UniProtKB for curated and automatically annotated protein entries
- UniRef for clustered sets of sequences
- UniParc for comprehensive archive of protein sequences
- Offers tools for protein sequence analysis and classification
- Supports proteomics research and functional genomics studies
- Facilitates cross-referencing with other biological databases
Protein Data Bank (PDB)
- Primary repository for three-dimensional structures of biological macromolecules
- Contains experimentally determined structures of proteins, nucleic acids, and complexes
- Crucial for structural biology and drug design research
- Features of PDB include:
- Atomic coordinates and related information for each structure
- Tools for structure visualization and analysis
- Links to related literature and functional annotations
- Supports various fields of research:
- Structural biology and protein folding studies
- Structure-based drug design and rational protein engineering
- Comparative structural genomics and evolutionary studies
Kyoto Encyclopedia of Genes and Genomes (KEGG)
- Integrated database resource for understanding high-level functions of biological systems
- Focuses on molecular interaction networks and biochemical pathways
- Combines genomic, chemical, and systemic functional information
- Key components of KEGG include:
- KEGG PATHWAY for metabolic and signaling pathway maps
- KEGG GENES for gene catalogs of sequenced genomes
- KEGG LIGAND for information on chemical compounds and reactions
- Supports systems biology and bioinformatics research:
- Facilitates interpretation of large-scale molecular datasets
- Enables pathway mapping and functional annotation of genes
- Supports studies on metabolic engineering and drug target identification