Allosteric regulation is a key mechanism for controlling enzyme activity in cells. It involves molecules binding to sites other than the active site, causing shape changes that affect enzyme function. This allows for quick, reversible adjustments to enzyme activity based on cellular needs.
Allosteric enzymes have unique structures and behaviors compared to non-allosteric enzymes. They often have multiple subunits, are more flexible, and show cooperative binding. This lets them respond sensitively to cellular signals and metabolic changes, helping maintain balance in important processes.
Allosteric Regulation of Enzyme Activity
Mechanism and Characteristics
- Allosteric regulation controls enzyme activity through effector molecules binding to sites other than the active site
- Allosteric enzymes contain regulatory sites separate from catalytic sites for fine-tuning activity
- Effector binding induces conformational changes altering substrate affinity or catalytic efficiency
- Provides rapid and reversible control to activate or inhibit enzyme activity
- Integrates cellular signals and metabolic status to modulate enzyme function
- Often exhibits cooperativity where binding of one effector influences subsequent molecule binding
- Crucial for cellular homeostasis and environmental adaptation (blood glucose regulation)
Structural Features
- Typically possess quaternary structures with multiple subunits (hemoglobin)
- Contain distinct regulatory sites separate from the active site
- Exhibit greater conformational flexibility allowing significant structural changes upon effector binding
- Active sites more susceptible to conformational changes induced by distant effector binding
- Intersubunit interfaces facilitate transmission of conformational changes throughout protein structure
Allosteric vs Non-allosteric Enzymes
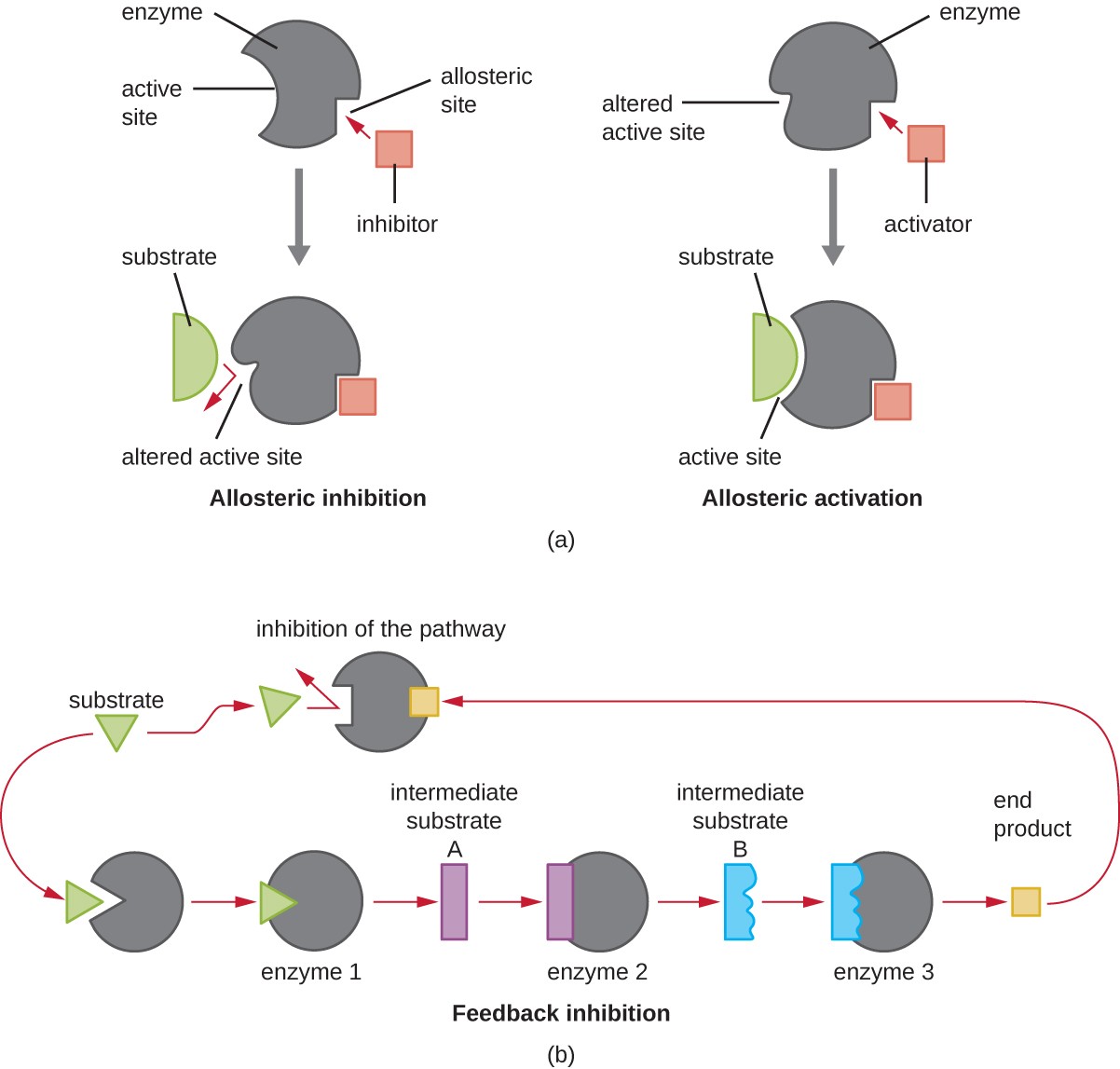
Structural Differences
- Allosteric enzymes often multi-subunit, non-allosteric may be monomeric or oligomeric
- Allosteric enzymes more conformationally flexible, non-allosteric maintain rigid structure
- Active sites of allosteric enzymes more influenced by binding at other locations
- Allosteric enzymes possess cooperative binding properties, typically absent in non-allosteric
- Non-allosteric enzymes have active sites less affected by events at other protein regions
Kinetic Differences
- Allosteric enzymes often display sigmoidal kinetics (oxygen binding to hemoglobin)
- Non-allosteric enzymes follow hyperbolic Michaelis-Menten kinetics (carbonic anhydrase)
- Hill coefficient quantifies cooperativity degree in allosteric enzymes
- Allosteric regulation alters enzyme's response to substrate concentration changes
- Advanced models analyze allosteric kinetics (Monod-Wyman-Changeux, Koshland-Némethy-Filmer)
Allosteric Activators and Inhibitors

Effects on Enzyme Kinetics
- Activators increase substrate affinity, shifting substrate-velocity curve leftward
- Inhibitors decrease substrate affinity, shifting substrate-velocity curve rightward
- Effectors alter Vmax and/or Km, changing catalytic efficiency
- Allosteric regulation often results in sigmoidal kinetics
- Hill coefficient describes sigmoidal curve steepness in allosteric enzymes
- Effectors influence enzyme response to substrate concentration changes
- Kinetic effects analyzed using advanced models (MWC, KNF)
Examples and Applications
- Fructose-2,6-bisphosphate activates phosphofructokinase-1 in glycolysis
- ATP inhibits phosphofructokinase-1, slowing glycolysis when energy levels high
- Oxygen acts as allosteric effector in hemoglobin, modulating oxygen binding affinity
- BPG (2,3-bisphosphoglycerate) allosterically regulates hemoglobin oxygen affinity in red blood cells
- Aspartate and CTP inhibit aspartate transcarbamoylase in pyrimidine synthesis
Physiological Significance of Allosteric Regulation
Metabolic Control
- Enables rapid, sensitive control of metabolic flux responding to cellular needs (glycolysis regulation)
- Facilitates feedback inhibition preventing overproduction (cholesterol synthesis)
- Allows feed-forward activation enhancing pathway efficiency (glycogen synthesis)
- Integrates multiple allosteric signals for complex regulation of metabolic pathways
- Maintains metabolic homeostasis balancing anabolic and catabolic processes
- Coordinates different metabolic pathways ensuring efficient resource use (Krebs cycle)
- Provides quick adaptation to changing energy demands without altering enzyme expression
Physiological Examples
- Phosphofructokinase-1 regulation controls glycolytic flux based on energy status
- Aspartate transcarbamoylase regulation balances pyrimidine nucleotide synthesis
- Hemoglobin's oxygen binding modulation adapts to tissue oxygen demands
- Glutamine synthetase regulation controls nitrogen metabolism in bacteria
- Glycogen phosphorylase regulation responds to glucose availability and hormonal signals