Human Genome Project Goals and Achievements
The Human Genome Project (HGP) was one of the largest coordinated biological research efforts in history. Its goal was to identify every gene in human DNA and sequence all 3 billion base pairs that make up the human genome. Understanding this project and its ripple effects is central to grasping how modern biology, medicine, and bioethics developed in the late 20th and early 21st centuries.
Project Overview and Timeline
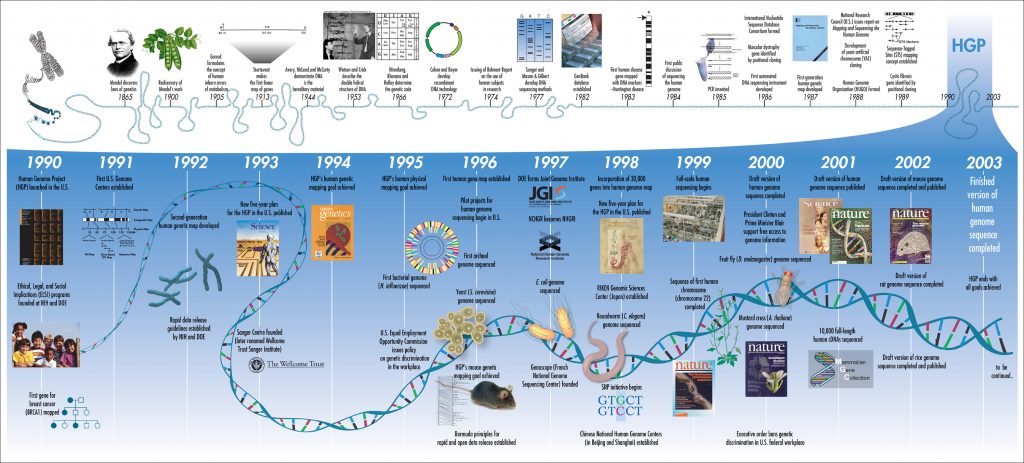
The HGP launched in 1990 as an international collaboration led by the U.S. National Institutes of Health and the Department of Energy, with contributions from research centers in the UK, France, Germany, Japan, and China. It was declared complete in April 2003, two years ahead of its original 15-year schedule and under budget.
A key detail for context: a privately funded parallel effort led by Craig Venter and Celera Genomics began in 1998 using a different sequencing strategy called "whole-genome shotgun" sequencing. The competition between the public HGP and Celera accelerated the timeline considerably. Both groups published draft sequences simultaneously in February 2001, and the final, high-quality reference sequence came from the public project in 2003.
Key Goals and Accomplishments
The project set out to give researchers the tools to understand genetic factors in human disease and to open new paths for diagnosis, treatment, and prevention. Its major accomplishments include:
- Sequencing approximately 3 billion base pairs of human DNA
- Identifying roughly 20,000–25,000 protein-coding genes (far fewer than the 100,000+ originally predicted, which itself was a significant finding about genome complexity)
- Mapping gene locations across all 23 pairs of human chromosomes
- Sequencing the genomes of key model organisms (mouse, fruit fly Drosophila, the roundworm C. elegans, yeast, and E. coli) to enable cross-species comparisons
That gene count surprise deserves emphasis. Scientists expected the complexity of the human body to require a huge number of genes. Finding only ~20,000–25,000 showed that much of biological complexity comes not from the number of genes, but from how genes are regulated, spliced, and expressed in different combinations.
Technological Advancements
The HGP didn't just produce data; it drove the development of technologies that transformed biological research:
- Automated DNA sequencing sped up the process of reading base pairs by orders of magnitude compared to earlier manual methods
- Bioinformatics emerged as a discipline to manage, store, and analyze the massive datasets the project generated
- Laboratory automation reduced costs and human error in sequencing workflows
These tools laid the groundwork for the dramatic drop in sequencing costs that followed. Sequencing a full human genome cost roughly $2.7 billion during the HGP era. By the 2020s, it costs under $1,000. That cost reduction is one of the most striking technology curves in modern science, outpacing even Moore's Law in computing.
Sequencing the Human Genome: Significance and Applications
Understanding Human Biology and Disease
The completed genome provides a reference map of the genetic information needed to build and maintain a human body. With this map, researchers can pinpoint genes involved in both common diseases (cancer, heart disease, diabetes) and rare genetic disorders (cystic fibrosis, Huntington's disease).
Before the HGP, finding a single disease-related gene could take years of painstaking work. After the project, genome-wide association studies (GWAS) made it possible to scan the entire genome for variants linked to disease, dramatically accelerating discovery. GWAS compare the genomes of large groups of people with and without a particular condition to find genetic variants that show up more frequently in the affected group.
Personalized Medicine and Drug Development
The genome revealed that humans share about 99.9% of their DNA. That remaining 0.1% of variation, however, can influence disease risk and drug response. These differences are the basis for personalized medicine, where treatments are matched to a patient's genetic profile rather than applied as one-size-fits-all.
For drug development, genomic data helps identify new drug targets (specific proteins or pathways involved in disease) and predict which patients are likely to benefit from or react badly to a given medication.
Comparative Genomics and Evolutionary Insights
Comparing the human genome to those of other species reveals conserved sequences: stretches of DNA that have remained largely unchanged across millions of years of evolution. If a sequence is conserved across distantly related species, it likely serves a critical biological function, because mutations in that region were selected against over time. These comparisons also clarify evolutionary relationships between species and help identify genes whose functions were previously unknown.
Increasing Accessibility
Advances in sequencing technology since 2003 have made genomic analysis far more accessible. What once required massive international infrastructure can now be done in individual hospital labs. This accessibility has opened the door to routine clinical use, from newborn screening panels to tumor profiling for cancer patients.
Ethical, Legal, and Social Issues of the Human Genome Project
The HGP was unusual among scientific projects in that it dedicated a portion of its budget (about 3–5%) specifically to studying the ethical, legal, and social implications (ELSI) of genomic research. This was a deliberate acknowledgment that mapping the human genome would raise questions science alone couldn't answer. The ELSI program became a model for how large-scale science projects could build ethical reflection into the research process from the start.
Genetic Privacy and Discrimination
Once your genome can be read, who gets to see it? The HGP raised serious concerns about genetic discrimination, where insurers, employers, or others might use genetic information against individuals. In the U.S., the Genetic Information Nondiscrimination Act (GINA), passed in 2008, was a direct legislative response. GINA prohibits health insurers and employers from discriminating based on genetic information, though it does not cover life insurance, disability insurance, or long-term care insurance. That gap matters: someone who tests positive for a high-risk mutation could still face consequences when applying for life insurance.
Beyond discrimination, there are broader ethical questions about using genetic information for non-medical purposes, such as selecting embryos for specific traits or making reproductive decisions based on carrier status.
Commercialization and Intellectual Property
Can a company patent a human gene? This question became a major legal battle. In 2013, the U.S. Supreme Court ruled in Association for Molecular Pathology v. Myriad Genetics that naturally occurring DNA sequences cannot be patented, though synthetically created complementary DNA (cDNA) can be. The case centered on Myriad's patents on the BRCA1 and BRCA2 genes linked to breast and ovarian cancer, which had allowed the company to control (and charge high prices for) diagnostic testing.
The broader tension remains: genomic data has enormous commercial value, but restricting access to it can slow research and limit who benefits. The HGP's own commitment to making sequence data publicly available (through the "Bermuda Principles" adopted in 1996, which required data to be released within 24 hours of generation) set an important precedent for open science.
Psychological and Social Implications
Learning you carry a gene variant associated with a serious disease (like the APOE4 variant linked to Alzheimer's risk) can cause significant anxiety, even when the information is probabilistic rather than deterministic. Not everyone who carries a risk variant will develop the disease. This distinction between risk and diagnosis is one of the hardest things for patients to navigate.
There are also equity concerns. Access to genetic testing and the personalized treatments it enables is not evenly distributed. Socioeconomic status, geography, and insurance coverage all affect who benefits from genomic advances. Additionally, the original HGP reference genome was built primarily from donors of European descent, which means it may not capture genetic variation in other populations as well. Efforts like the All of Us Research Program are working to address this gap.

Informed Consent and Research Ethics
Genomic research requires careful handling of informed consent. Participants need to understand how their genetic data will be used, stored, and shared. This is complicated by the fact that genetic information is inherently familial: sequencing one person reveals information about their relatives, who may not have consented.
The rise of direct-to-consumer genetic testing (companies like 23andMe) adds another layer. Consumers may receive complex risk information without the guidance of a genetic counselor, leading to potential misinterpretation. A result showing "elevated risk" for a condition is not the same as a diagnosis, but that distinction can be lost without proper context.
Personalized Medicine and its Impact on Healthcare
Tailoring Prevention, Diagnosis, and Treatment
Personalized medicine (also called precision medicine) uses an individual's genetic information to guide healthcare decisions. Rather than treating all patients with the same condition identically, clinicians can factor in genetic variations that affect disease risk, progression, and treatment response.
Genetic testing can identify people who carry mutations for inherited disorders (like BRCA mutations for breast cancer) before symptoms appear. This allows for earlier interventions such as increased surveillance, preventive medications, or even prophylactic surgery.
Pharmacogenomics and Targeted Therapies
Pharmacogenomics studies how a person's genes affect their response to drugs. Some practical examples:
- Variants in the CYP2D6 gene affect how patients metabolize codeine. Poor metabolizers get little pain relief; ultra-rapid metabolizers can experience dangerous toxicity from standard doses.
- The drug warfarin (a blood thinner) requires careful dosing that varies significantly based on variants in the CYP2C9 and VKORC1 genes. Without genetic testing, doctors rely on trial-and-error dose adjustments, which can lead to dangerous bleeding or clotting.
- In cancer treatment, drugs like trastuzumab (Herceptin) target tumors that overexpress the HER2 protein. Patients whose tumors lack this marker won't benefit from the drug, so genetic testing of the tumor guides treatment selection.
These examples show how pharmacogenomics reduces adverse drug reactions and avoids prescribing medications that won't work for a given patient.
Shifting Focus to Proactive Prevention
One of the most significant promises of personalized medicine is moving healthcare from a reactive model (treating disease after it appears) to a proactive one (identifying risk and intervening early). If genetic screening reveals that a patient has elevated risk for colorectal cancer, for instance, earlier and more frequent colonoscopies can catch problems at a treatable stage.
Integration of Genomic and Clinical Data
Genomic information is most useful when combined with other clinical data: family history, lifestyle factors, environmental exposures, and lab results. No gene operates in isolation. A genetic predisposition to type 2 diabetes, for example, interacts with diet, exercise, and other factors to determine actual disease risk. This is why personalized medicine is better understood as genomics-informed medicine rather than genomics-determined medicine.
Improving Healthcare Efficiency
Personalized approaches have the potential to reduce healthcare costs by:
- Avoiding trial-and-error prescribing (getting the right drug to the right patient faster)
- Reducing hospitalizations from adverse drug reactions
- Focusing expensive screening and prevention resources on patients who will benefit most
The challenge is implementation. Integrating genomic data into routine clinical workflows requires training clinicians, updating electronic health records, and ensuring equitable access across different healthcare settings. As of the 2020s, pharmacogenomic testing is becoming more common but is still far from standard practice in most clinical environments.