RNA Splicing
Pre-mRNA Processing and Splicing
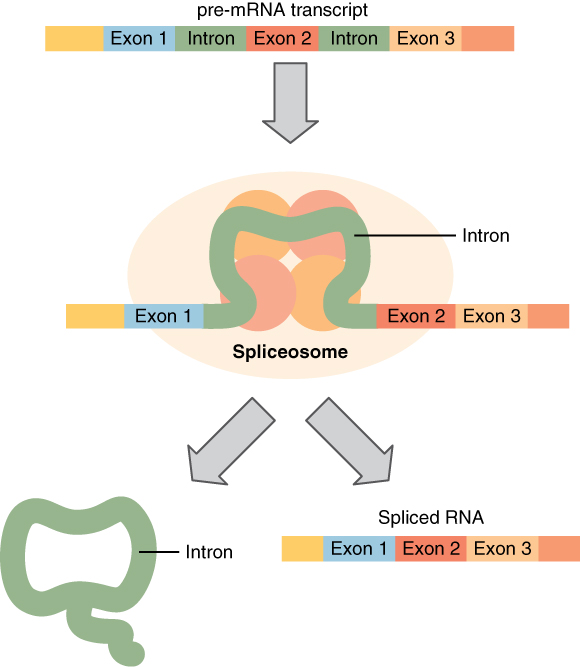
Pre-mRNA is the initial transcript produced by RNA polymerase II. It contains both exons (coding regions) and introns (non-coding intervening sequences). Before the mRNA can be exported to the cytoplasm for translation, introns must be removed through a process called splicing, which excises the introns and joins the remaining exons into a continuous, mature mRNA molecule.
Spliceosome and Alternative Splicing
Splicing is carried out by a large molecular machine called the spliceosome, composed of five small nuclear ribonucleoproteins (snRNPs, pronounced "snurps": U1, U2, U4, U5, and U6) along with numerous associated proteins. The spliceosome recognizes conserved sequences at exon-intron boundaries, known as splice sites: the 5' splice site (GU), the 3' splice site (AG), and the branch point (an adenosine residue upstream of the 3' splice site).
The splicing reaction proceeds in two transesterification steps:
- The 2'-OH of the branch point adenosine attacks the phosphodiester bond at the 5' splice site, forming a lariat intermediate (a looped intron structure) and freeing the upstream exon.
- The free 3'-OH of the upstream exon attacks the phosphodiester bond at the 3' splice site, joining the two exons and releasing the lariat intron.
Alternative splicing allows a single gene to produce multiple mRNA variants by including or excluding different combinations of exons. The CD44 gene is a dramatic example: through alternative splicing, it can generate over 100 distinct protein isoforms from one pre-mRNA. This is a major source of proteomic diversity in eukaryotes.
mRNA Modifications
5' Capping and 3' Polyadenylation
Mature mRNA is modified at both ends before export from the nucleus.
5' capping occurs co-transcriptionally (while the transcript is still being made) and involves the addition of a 7-methylguanosine (m7G) cap to the 5' end via an unusual 5'-to-5' triphosphate linkage. This cap:
- Protects the mRNA from exonuclease degradation
- Recruits the ribosome and translation initiation factors (especially eIF4E)
- Facilitates splicing of the first intron and nuclear export
3' polyadenylation adds a poly(A) tail, typically 100–250 adenine nucleotides, to the 3' end. The process requires recognition of a polyadenylation signal (usually AAUAAA) in the pre-mRNA, followed by cleavage and addition of the tail by poly(A) polymerase (PAP). The poly(A) tail:
- Enhances mRNA stability by slowing 3' exonuclease degradation
- Facilitates nuclear export
- Promotes translation initiation (through interactions between poly(A)-binding protein and eIF4G)

RNA Editing
RNA editing alters the nucleotide sequence of an RNA molecule after transcription, without changing the DNA template. This can involve insertion, deletion, or chemical modification of specific bases.
The most common form in mammals is adenosine-to-inosine (A-to-I) editing, catalyzed by ADAR enzymes. Because inosine is read as guanosine by the translational machinery, this effectively changes an A to a G in the coding message.
A classic example is the apolipoprotein B (apoB) gene. In the liver, the full-length mRNA is translated into apoB-100 (a 4,536-amino-acid protein involved in LDL cholesterol transport). In the intestine, a cytidine deaminase converts a specific C to U, creating a premature stop codon. This produces the shorter apoB-48 isoform, which functions in dietary lipid absorption. One gene, two functionally distinct proteins, generated entirely through RNA editing.
Non-coding RNA Processing
tRNA and rRNA Processing
Non-coding RNAs require their own processing pathways to become functional.
tRNA processing involves several steps:
- The 5' leader sequence is removed by RNase P (a ribozyme).
- The 3' trailer is trimmed by exonucleases.
- The universal CCA trinucleotide is added to the 3' end by tRNA nucleotidyltransferase (this is where the amino acid will attach).
- Specific bases are chemically modified (e.g., pseudouridine, dihydrouridine, inosine). Modifications in the anticodon loop are particularly important because they fine-tune codon-anticodon recognition and translational accuracy.
- Some tRNAs contain introns that are removed by a dedicated splicing endonuclease (distinct from the spliceosome).
rRNA processing starts with a large polycistronic precursor (47S in humans), transcribed by RNA polymerase I, that contains the 18S, 5.8S, and 28S rRNA sequences separated by internal and external transcribed spacers. Processing involves:
- Cleavage of spacer regions by endo- and exonucleases
- Extensive chemical modifications, primarily 2'-O-methylation and pseudouridylation, guided by small nucleolar RNAs (snoRNAs) through base-pairing with the rRNA target
- Assembly with ribosomal proteins in the nucleolus to form the 40S and 60S ribosomal subunits
The 5S rRNA is transcribed separately by RNA polymerase III and joins the large subunit during ribosome assembly.