Enzymes are protein catalysts that accelerate chemical reactions in living organisms. They're central to metabolism, digestion, and cellular signaling, and because of their specificity, they make excellent drug targets. Understanding how enzymes work, how they're inhibited, and how they're regulated is foundational to medicinal chemistry and rational drug design.
Enzymes as biological catalysts
Enzymes lower the activation energy of reactions, allowing them to proceed much faster than they would spontaneously. Without enzymes, most biochemical reactions would be far too slow to sustain life.
- They are highly specific, typically catalyzing only one reaction or a narrow set of related reactions
- They aren't consumed in the reaction and can be used repeatedly
- Their activity can be tightly regulated, making them ideal control points in metabolic pathways and, by extension, ideal targets for drugs
Structure of enzymes
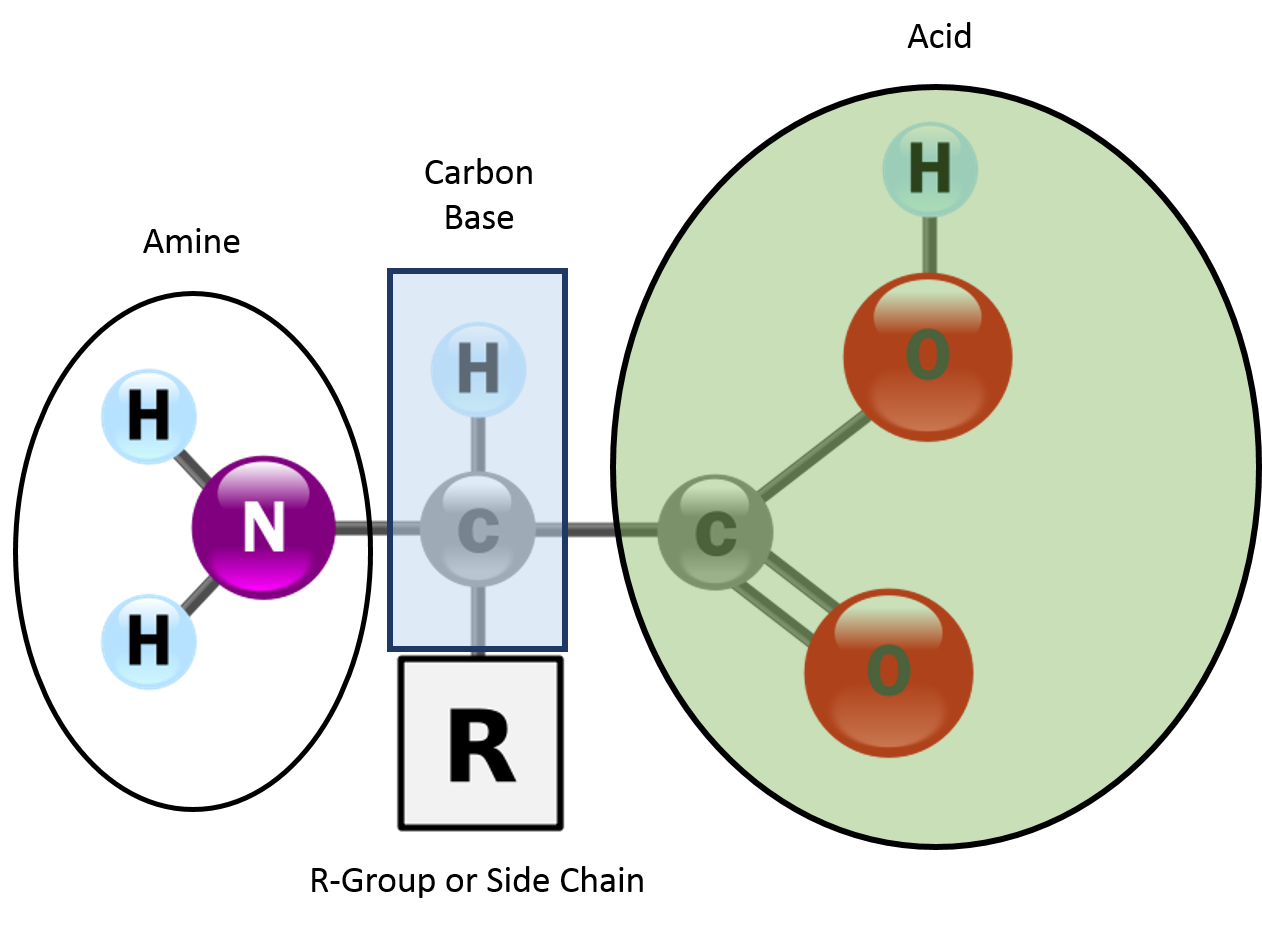
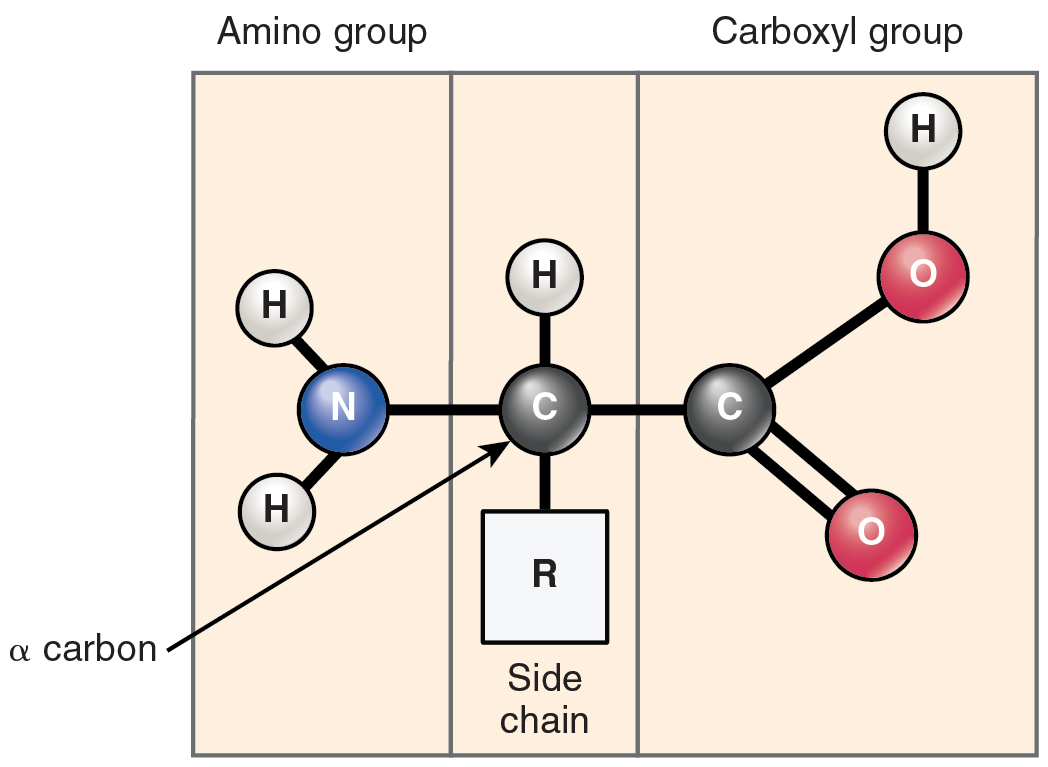
Amino acid composition
Enzymes are built from amino acids linked by peptide bonds. The specific sequence of amino acids (encoded by genes) determines each enzyme's unique shape and catalytic behavior. Even small changes in amino acid composition can affect stability, solubility, and how well the enzyme catalyzes its reaction.
Primary, secondary, tertiary structure
- Primary structure is the linear sequence of amino acids in the polypeptide chain. Think of it as the "spelling" of the protein.
- Secondary structure refers to local folding patterns, mainly α-helices and β-sheets, held together by hydrogen bonds between backbone atoms.
- Tertiary structure is the overall 3D shape of the enzyme, formed by interactions between amino acid side chains (R-groups). These include disulfide bonds, hydrophobic interactions, ionic bonds, and hydrogen bonds.
The tertiary structure is what creates the active site, so it directly determines catalytic function.
Active site for substrate binding
The active site is a specific cleft or pocket on the enzyme surface where the substrate binds and the reaction takes place. It's formed by the enzyme's tertiary (and sometimes quaternary) structure, bringing together amino acid residues that may be far apart in the primary sequence.
These residues interact with the substrate through complementary charge, shape, and hydrophobicity, positioning it precisely for catalysis.
Enzyme-substrate interactions
Lock and key model
This older model proposes that the active site has a rigid shape perfectly complementary to the substrate, like a key fitting into a lock. It explains enzyme specificity well but fails to account for the flexibility that enzymes actually display during catalysis.
Induced fit model
The induced fit model is the more accepted explanation. Here, the active site is flexible and adjusts its shape when the substrate binds. The initial enzyme-substrate contact triggers a conformational change that optimizes the fit and stabilizes the transition state. This model explains why some enzymes can accommodate slightly different substrates and why binding itself can contribute to catalysis.
Substrate specificity of enzymes
Enzymes are highly selective about which substrates they act on. This specificity comes from the precise geometry, size, and chemical environment of the active site. It ensures that each enzyme catalyzes only its intended reaction, preventing unwanted interference with other cellular processes.
Enzyme kinetics
Factors affecting reaction rates
- Temperature: Higher temperatures increase molecular motion and reaction rates, but only up to a point. Beyond the enzyme's optimal temperature, it denatures (loses its 3D shape) and activity drops sharply.
- pH: Each enzyme has an optimal pH range. Extreme pH values alter the ionization of active site residues and can denature the enzyme. For example, pepsin works best at pH ~2 (stomach), while trypsin prefers pH ~8 (small intestine).
- Substrate concentration: Increasing substrate concentration raises the reaction rate, but only until all enzyme molecules are occupied. At that point, the enzyme is saturated and the rate plateaus.
Michaelis-Menten equation
The Michaelis-Menten equation models how reaction velocity relates to substrate concentration under steady-state conditions:
This equation assumes a single substrate and that the enzyme-substrate complex forms and breaks down at steady state. It's the starting point for characterizing enzyme kinetics and extracting key parameters.
Km and Vmax parameters
- (Michaelis constant) is the substrate concentration at which the reaction rate equals half of . A lower means the enzyme reaches half-maximal speed at a lower substrate concentration, indicating higher substrate affinity.
- is the maximum rate when every enzyme molecule is bound to substrate. It depends on both enzyme concentration and the enzyme's intrinsic catalytic rate constant ().
A useful measure of catalytic efficiency is , which accounts for both how fast the enzyme works and how well it binds substrate.
Enzyme inhibition
Competitive vs noncompetitive inhibition
Competitive inhibitors bind to the active site, directly competing with the substrate. They typically resemble the substrate structurally. In kinetic terms, competitive inhibition increases the apparent (the enzyme appears to have lower affinity for substrate) but does not change , because adding enough substrate can outcompete the inhibitor.
Noncompetitive inhibitors bind at a site other than the active site (an allosteric site), causing a conformational change that reduces catalytic activity. They decrease but leave unchanged, since the inhibitor doesn't interfere with substrate binding directly.
Reversible vs irreversible inhibitors
- Reversible inhibitors bind non-covalently and can dissociate from the enzyme. They include competitive, noncompetitive, and uncompetitive types. Their effects can be overcome by removing the inhibitor (e.g., by dilution or dialysis).
- Irreversible inhibitors form covalent bonds with the enzyme, permanently inactivating it. Many clinically important drugs work this way. For example, aspirin irreversibly acetylates cyclooxygenase (COX), blocking prostaglandin synthesis.
Inhibition constants (Ki)
The inhibition constant () quantifies an inhibitor's potency. It represents the inhibitor concentration needed to achieve half-maximal inhibition. A lower means a more potent inhibitor, requiring less drug to produce the same effect. In drug design, values help compare candidate molecules and guide optimization.
Regulation of enzyme activity
Allosteric regulation
Allosteric regulation occurs when effector molecules bind to sites distinct from the active site (allosteric sites). Binding of an allosteric activator shifts the enzyme toward a more active conformation, while an allosteric inhibitor shifts it toward a less active form.
This mechanism allows cells to fine-tune enzyme activity in response to changing conditions. Many allosteric enzymes display sigmoidal (S-shaped) kinetics rather than the hyperbolic curve predicted by Michaelis-Menten.
Covalent modifications
Enzymes can be switched on or off through covalent modifications such as:
- Phosphorylation (addition of a phosphate group by kinases; removal by phosphatases)
- Acetylation (addition of an acetyl group)
- Glycosylation (addition of sugar moieties)
These modifications alter the enzyme's charge, shape, or interactions, changing its activity, stability, or localization. Phosphorylation is especially common in signal transduction pathways and is a frequent target in drug development.
Zymogen activation
Zymogens (proenzymes) are inactive enzyme precursors that require proteolytic cleavage to become active. This is a safety mechanism: the enzyme is synthesized in an inactive form and activated only where and when it's needed.
- Pepsinogen → pepsin (in the stomach)
- Trypsinogen → trypsin (in the small intestine)
- Prothrombin → thrombin (in the blood clotting cascade)
This prevents premature digestion of tissues or inappropriate clotting.
Coenzymes and cofactors
Role in enzyme catalysis
Many enzymes require non-protein helpers to function. Cofactors are inorganic ions (e.g., , , ), while coenzymes are organic molecules, often derived from vitamins. Together, they assist with electron transfer, group transfer, or stabilizing the transition state.
An enzyme without its required cofactor/coenzyme is called an apoenzyme. The complete, catalytically active form (enzyme + cofactor) is the holoenzyme.
Common examples (NAD+, FAD, etc.)
- / : Derived from niacin (vitamin B3). Act as electron carriers in redox reactions, shuttling hydride ions between metabolic pathways.
- : Derived from riboflavin (vitamin B2). Another redox coenzyme, tightly bound to enzymes like succinate dehydrogenase.
- Coenzyme A (CoA): Derived from pantothenic acid (vitamin B5). Carries acyl groups and is essential in the citric acid cycle and fatty acid metabolism.
Vitamin-derived coenzymes
- Thiamine pyrophosphate (TPP) from vitamin B1: cofactor for decarboxylation reactions (e.g., pyruvate dehydrogenase)
- Pyridoxal phosphate (PLP) from vitamin B6: involved in transamination and other amino acid transformations
- Biotin (vitamin B7): carries carboxyl groups in carboxylation reactions (e.g., pyruvate carboxylase)
This vitamin-coenzyme connection is why vitamin deficiencies can cause metabolic disorders.
Enzyme classification
International enzyme nomenclature
The International Union of Biochemistry and Molecular Biology (IUBMB) maintains a standardized naming system. Enzymes are named based on the reaction they catalyze, with the suffix "-ase" appended to the substrate name or reaction type (e.g., lactase hydrolyzes lactose, kinase transfers phosphate groups).
Six main enzyme classes
| Class | Reaction Type | Example |
|---|---|---|
| Oxidoreductases | Redox reactions (electron transfer) | Alcohol dehydrogenase |
| Transferases | Transfer of functional groups (methyl, phosphoryl, acyl) | Hexokinase |
| Hydrolases | Hydrolysis of bonds (esters, peptides, glycosides) | Lipase |
| Lyases | Non-hydrolytic removal/addition of groups, often forming double bonds | Fumarase |
| Isomerases | Rearrangement of atoms within a molecule | Phosphoglucose isomerase |
| Ligases | Joining two molecules, coupled with ATP hydrolysis | DNA ligase |
EC numbering system
Each enzyme receives a four-digit EC number in the format EC x.y.z.w:
- First digit (x): Main class (1 through 6, corresponding to the six classes above)
- Second digit (y): Subclass (type of bond or group involved)
- Third digit (z): Sub-subclass (more specific reaction detail)
- Fourth digit (w): Serial number of the specific enzyme
For example, EC 1.1.1.1 is alcohol dehydrogenase (class 1 = oxidoreductase).
Enzymes as drug targets
Rational drug design strategies
Enzymes involved in disease pathways are prime drug targets because their active sites offer well-defined binding pockets.
- Structure-based drug design uses X-ray crystallography or cryo-EM data of the enzyme's 3D structure to design molecules that fit the active site and block function.
- Ligand-based drug design starts from known inhibitors or substrates and modifies their structure to improve potency, selectivity, or pharmacokinetic properties.
Both approaches aim to create molecules with high affinity for the target enzyme and minimal off-target effects.
Examples of enzyme-targeted drugs
- Statins (atorvastatin, simvastatin): Competitive inhibitors of HMG-CoA reductase, the rate-limiting enzyme in cholesterol biosynthesis. Used to treat hypercholesterolemia.
- ACE inhibitors (captopril, enalapril): Block angiotensin-converting enzyme, preventing conversion of angiotensin I to the vasoconstrictor angiotensin II. Used for hypertension and heart failure.
- HIV protease inhibitors (saquinavir, ritonavir): Target viral proteases essential for processing HIV polyproteins into functional viral components.
Advantages and challenges
Advantages:
- High specificity allows for potent drugs with fewer off-target side effects
- Well-characterized active sites make rational design feasible
- Kinetic parameters (, ) provide clear metrics for drug optimization
Challenges:
- Drug resistance can develop, especially with rapidly mutating targets like viral enzymes
- Enzyme redundancy in cells may provide compensatory pathways that reduce drug effectiveness
- Selectivity between closely related enzyme isoforms can be difficult to achieve
Enzyme immobilization
Methods for enzyme immobilization
Immobilization attaches enzymes to solid supports or confines them within matrices, making them easier to handle in industrial and clinical settings.
- Adsorption: Enzyme binds to a support surface through weak interactions (van der Waals, ionic). Simple but can lead to enzyme leaching.
- Covalent bonding: Enzyme is chemically linked to the support via covalent bonds. Strong attachment, but the chemistry must avoid disrupting the active site.
- Entrapment: Enzyme is physically trapped within a polymer gel or membrane. The enzyme stays free in solution within the matrix but can't escape.
- Cross-linking: Enzymes are linked to each other (or to a support) using bifunctional reagents like glutaraldehyde. No separate support needed, but can reduce activity.
Applications in industry and medicine
- High-fructose corn syrup production uses immobilized glucose isomerase to convert glucose to fructose
- Lactose-free milk is produced using immobilized lactase to break down lactose
- Biosensors (e.g., glucose monitors for diabetic patients) use immobilized enzymes to detect specific analytes
- Bioreactors with immobilized enzymes treat industrial waste and synthesize pharmaceuticals
Improved stability and reusability
Immobilization protects enzymes from denaturation and aggregation, extending their functional lifespan. The enzyme can be easily separated from the product and reused across many reaction cycles. This reusability dramatically reduces costs, making enzymatic processes economically viable at industrial scale.
Enzymes in disease and diagnosis
Enzyme deficiencies and disorders
Inborn errors of metabolism are genetic conditions caused by mutations that reduce or eliminate a specific enzyme's activity. Without the enzyme, substrates accumulate to toxic levels or essential products aren't made.
- Phenylketonuria (PKU): Deficiency in phenylalanine hydroxylase leads to toxic buildup of phenylalanine, causing intellectual disability if untreated.
- Galactosemia: Deficiency in galactose-1-phosphate uridyltransferase prevents proper galactose metabolism, leading to liver damage and developmental issues.
Diagnostic enzyme assays
Measuring enzyme levels in blood or other body fluids helps diagnose tissue damage and disease:
- ALT and AST (alanine and aspartate aminotransferases): Elevated levels indicate liver damage or hepatitis
- Creatine kinase (CK): Elevated after myocardial infarction (heart attack) or skeletal muscle injury. CK-MB is the isoform most specific to cardiac tissue.
- Alkaline phosphatase (ALP): Elevated in bone disorders and biliary obstruction
The principle is straightforward: when cells are damaged, intracellular enzymes leak into the bloodstream, and their elevated levels serve as biomarkers.
Enzyme replacement therapy
Enzyme replacement therapy (ERT) delivers a functional recombinant version of the missing enzyme to patients with genetic enzyme deficiencies.
- Gaucher disease: Treated with recombinant glucocerebrosidase (imiglucerase)
- Fabry disease: Treated with recombinant -galactosidase A (agalsidase)
ERT can significantly improve quality of life, but it has limitations: the recombinant enzyme may trigger immune responses, treatment requires repeated intravenous infusions, and production costs are very high. Delivery to certain tissues (especially the brain, due to the blood-brain barrier) remains a major challenge.